The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
survalis provides a unified framework for survival
machine learning survival analysis in R. It supports a wide range of
learners, evaluation metrics, cross-validation and interpretability
methods.
# Install from GitHub
remotes::install_github("ielbadisy/survalis")
# Or from source
install.packages("survalis_0.7.0.tar.gz", repos = NULL, type = "source")fit_*(),
predict_*(), tune_*()mlsurv_model objectsdata.frame of survival
probabilities: t=100, t=200, …fit_*/predict_* with
cv_survlearner() or score_survmodel()library(survalis)
# See all available learners
list_survlearners()
#> # A tibble: 19 × 8
#> learner fit predict tune has_fit has_predict has_tune available
#> <chr> <chr> <chr> <chr> <lgl> <lgl> <lgl> <lgl>
#> 1 coxph fit_cox… predic… <NA> TRUE TRUE FALSE TRUE
#> 2 aalen fit_aal… predic… <NA> TRUE TRUE FALSE TRUE
#> 3 glmnet fit_glm… predic… tune… TRUE TRUE TRUE TRUE
#> 4 selectcox fit_sel… predic… tune… TRUE TRUE TRUE TRUE
#> 5 aftgee fit_aft… predic… <NA> TRUE TRUE FALSE TRUE
#> 6 flexsurvreg fit_fle… predic… tune… TRUE TRUE TRUE TRUE
#> 7 stpm2 fit_stp… predic… <NA> TRUE TRUE FALSE TRUE
#> 8 bnnsurv fit_bnn… predic… tune… TRUE TRUE TRUE TRUE
#> 9 rpart fit_rpa… predic… tune… TRUE TRUE TRUE TRUE
#> 10 bart fit_bart predic… tune… TRUE TRUE TRUE TRUE
#> 11 xgboost fit_xgb… predic… tune… TRUE TRUE TRUE TRUE
#> 12 ranger fit_ran… predic… tune… TRUE TRUE TRUE TRUE
#> 13 rsf fit_rsf predic… tune… TRUE TRUE TRUE TRUE
#> 14 cforest fit_cfo… predic… tune… TRUE TRUE TRUE TRUE
#> 15 blackboost fit_bla… predic… tune… TRUE TRUE TRUE TRUE
#> 16 survsvm fit_sur… predic… tune… TRUE TRUE TRUE TRUE
#> 17 survdnn fit_sur… predic… tune… TRUE TRUE TRUE TRUE
#> 18 orsf fit_orsf predic… tune… TRUE TRUE TRUE TRUE
#> 19 survmetalearner fit_sur… predic… <NA> TRUE TRUE FALSE TRUE
# See only tunable learners (those with a tune_* function)
list_survlearners(has_tune = TRUE)
#> # A tibble: 14 × 8
#> learner fit predict tune has_fit has_predict has_tune available
#> <chr> <chr> <chr> <chr> <lgl> <lgl> <lgl> <lgl>
#> 1 glmnet fit_glmnet predic… tune… TRUE TRUE TRUE TRUE
#> 2 selectcox fit_selectc… predic… tune… TRUE TRUE TRUE TRUE
#> 3 flexsurvreg fit_flexsur… predic… tune… TRUE TRUE TRUE TRUE
#> 4 bnnsurv fit_bnnsurv predic… tune… TRUE TRUE TRUE TRUE
#> 5 rpart fit_rpart predic… tune… TRUE TRUE TRUE TRUE
#> 6 bart fit_bart predic… tune… TRUE TRUE TRUE TRUE
#> 7 xgboost fit_xgboost predic… tune… TRUE TRUE TRUE TRUE
#> 8 ranger fit_ranger predic… tune… TRUE TRUE TRUE TRUE
#> 9 rsf fit_rsf predic… tune… TRUE TRUE TRUE TRUE
#> 10 cforest fit_cforest predic… tune… TRUE TRUE TRUE TRUE
#> 11 blackboost fit_blackbo… predic… tune… TRUE TRUE TRUE TRUE
#> 12 survsvm fit_survsvm predic… tune… TRUE TRUE TRUE TRUE
#> 13 survdnn fit_survdnn predic… tune… TRUE TRUE TRUE TRUE
#> 14 orsf fit_orsf predic… tune… TRUE TRUE TRUE TRUE
# Shortcut for tunable learners
list_tunable_survlearners()
#> # A tibble: 14 × 8
#> learner fit predict tune has_fit has_predict has_tune available
#> <chr> <chr> <chr> <chr> <lgl> <lgl> <lgl> <lgl>
#> 1 glmnet fit_glmnet predic… tune… TRUE TRUE TRUE TRUE
#> 2 selectcox fit_selectc… predic… tune… TRUE TRUE TRUE TRUE
#> 3 flexsurvreg fit_flexsur… predic… tune… TRUE TRUE TRUE TRUE
#> 4 bnnsurv fit_bnnsurv predic… tune… TRUE TRUE TRUE TRUE
#> 5 rpart fit_rpart predic… tune… TRUE TRUE TRUE TRUE
#> 6 bart fit_bart predic… tune… TRUE TRUE TRUE TRUE
#> 7 xgboost fit_xgboost predic… tune… TRUE TRUE TRUE TRUE
#> 8 ranger fit_ranger predic… tune… TRUE TRUE TRUE TRUE
#> 9 rsf fit_rsf predic… tune… TRUE TRUE TRUE TRUE
#> 10 cforest fit_cforest predic… tune… TRUE TRUE TRUE TRUE
#> 11 blackboost fit_blackbo… predic… tune… TRUE TRUE TRUE TRUE
#> 12 survsvm fit_survsvm predic… tune… TRUE TRUE TRUE TRUE
#> 13 survdnn fit_survdnn predic… tune… TRUE TRUE TRUE TRUE
#> 14 orsf fit_orsf predic… tune… TRUE TRUE TRUE TRUE# List available interpretability methods
list_interpretability_methods()
#> # A tibble: 8 × 4
#> compute plot has_compute has_plot
#> <chr> <chr> <lgl> <lgl>
#> 1 compute_shap plot_shap TRUE TRUE
#> 2 compute_pdp plot_pdp TRUE TRUE
#> 3 compute_ale plot_ale TRUE TRUE
#> 4 compute_surrogate plot_surrogate TRUE TRUE
#> 5 compute_tree_surrogate plot_tree_surrogate TRUE TRUE
#> 6 compute_varimp plot_varimp TRUE TRUE
#> 7 compute_interactions plot_interactions TRUE TRUE
#> 8 compute_counterfactual <NA> TRUE FALSE
# Show which compute_* methods have a plot_* counterpart
subset(list_interpretability_methods(), !is.na(plot))
#> # A tibble: 7 × 4
#> compute plot has_compute has_plot
#> <chr> <chr> <lgl> <lgl>
#> 1 compute_shap plot_shap TRUE TRUE
#> 2 compute_pdp plot_pdp TRUE TRUE
#> 3 compute_ale plot_ale TRUE TRUE
#> 4 compute_surrogate plot_surrogate TRUE TRUE
#> 5 compute_tree_surrogate plot_tree_surrogate TRUE TRUE
#> 6 compute_varimp plot_varimp TRUE TRUE
#> 7 compute_interactions plot_interactions TRUE TRUE# List available metrics used in cross-validation and scoring
list_metrics()
#> # A tibble: 3 × 4
#> metric direction summary range
#> <chr> <chr> <chr> <chr>
#> 1 cindex maximize Harrell-style concordance index for survival predictio… [0, …
#> 2 brier minimize Brier Score at specified evaluation time(s) (IPCW-weig… [0, …
#> 3 ibs minimize Integrated Brier Score over an evaluation time grid (I… [0, …1. Fit a model
mod_cox <- fit_coxph(Surv(time, status) ~ age + karno + celltype, data = veteran)
summary(mod_cox)
#>
#> ── coxph summary ───────────────────────────────────────────────────────────────
#> Formula:
#> Surv(time, status) ~ age + karno + celltype
#> Engine: survival
#> Learner: coxph
#> Data summary:
#> - Observations: 137
#> - Predictors: "age, karno, celltypesmallcell, celltypeadeno, celltypelarge"
#> - Time range: [1, 999]
#> - Event rate: "93.4%"2. Predict survival probabilities
pred <- predict_coxph(mod_cox, newdata = veteran[1:5, ], times = c(100, 200))
pred
#> t=100 t=200
#> 1 0.6142681 0.3541697
#> 2 0.6944383 0.4599242
#> 3 0.5556797 0.2860796
#> 4 0.6033305 0.3408724
#> 5 0.6959633 0.46207833. Evaluate model performance
Direct evalution (single split):
score <- score_survmodel(mod_cox, times = c(100, 200), metrics = c("cindex", "ibs"))
score
#> # A tibble: 2 × 2
#> metric value
#> <chr> <dbl>
#> 1 cindex 0.734
#> 2 ibs 0.160cv_res <- cv_survlearner(
Surv(time, status) ~ age + karno + celltype,
veteran,
fit_coxph,
predict_coxph,
times = 80,
metrics = c("cindex", "ibs"),
folds = 5,
seed = 123,
verbose = FALSE
)
cv_res
#> # A tibble: 10 × 5
#> splits id fold metric value
#> <list> <chr> <int> <chr> <dbl>
#> 1 <split [109/28]> Fold1 1 cindex 0.699
#> 2 <split [109/28]> Fold1 1 ibs 0.227
#> 3 <split [109/28]> Fold2 2 cindex 0.812
#> 4 <split [109/28]> Fold2 2 ibs 0.141
#> 5 <split [110/27]> Fold3 3 cindex 0.695
#> 6 <split [110/27]> Fold3 3 ibs 0.217
#> 7 <split [110/27]> Fold4 4 cindex 0.698
#> 8 <split [110/27]> Fold4 4 ibs 0.188
#> 9 <split [110/27]> Fold5 5 cindex 0.688
#> 10 <split [110/27]> Fold5 5 ibs 0.138cv_summary(cv_res)
#> # A tibble: 2 × 7
#> metric mean sd n se lower upper
#> <chr> <dbl> <dbl> <int> <dbl> <dbl> <dbl>
#> 1 cindex 0.718 0.0528 5 0.0236 0.672 0.765
#> 2 ibs 0.182 0.0417 5 0.0186 0.146 0.2194. Visualize interpretation
shap_meanabs <- compute_shap(
model = mod_cox,
newdata = veteran[100,],
baseline_data = veteran,
times = 80,
sample.size = 50,
aggregate = TRUE,
method = "meanabs"
)
shap_meanabs
#> feature phi
#> trt trt 0.000000000
#> celltype celltype 0.005850095
#> karno karno 0.045326970
#> diagtime diagtime 0.000000000
#> age age 0.002540069
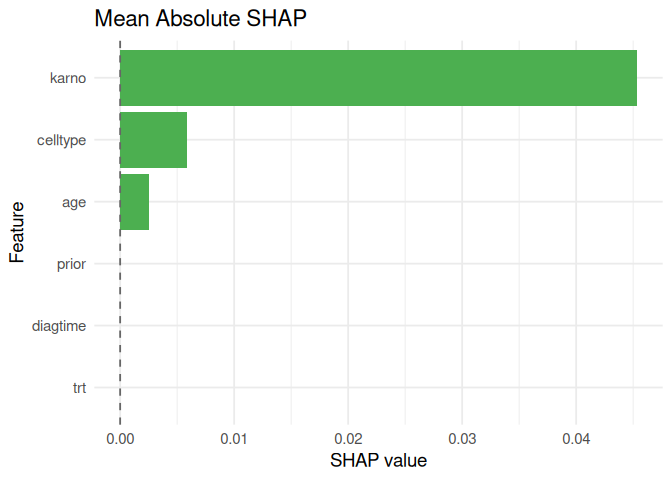
#> prior prior 0.000000000plot_shap(shap_meanabs)
survalis also provides PDP, ALE, surrogate explanations,
tree surrogates, permutation importance, interaction analysis, and
counterfactuals.
Partial dependence and ICE
pdp_age <- compute_pdp(
model = mod_cox,
data = veteran,
feature = "age",
times = c(100, 200, 300),
method = "pdp+ice"
)
plot_pdp(pdp_age, feature = "age", which = "per_time")
#> Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
#> ℹ Please use tidy evaluation idioms with `aes()`.
#> ℹ See also `vignette("ggplot2-in-packages")` for more information.
#> ℹ The deprecated feature was likely used in the survalis package.
#> Please report the issue to the authors.
#> This warning is displayed once per session.
#> Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
#> generated.
#> Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
#> ℹ Please use `linewidth` instead.
#> ℹ The deprecated feature was likely used in the survalis package.
#> Please report the issue to the authors.
#> This warning is displayed once per session.
#> Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
#> generated.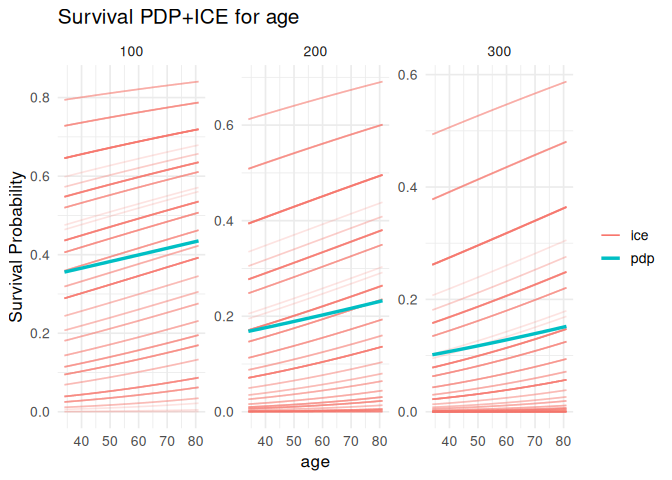
plot_pdp(pdp_age, feature = "age", which = "integrated", smooth = TRUE)
#> `geom_smooth()` using formula = 'y ~ x'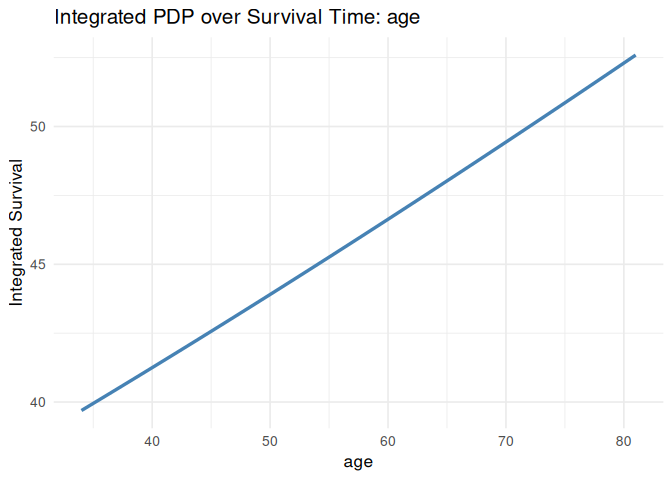
Accumulated local effects
ale_karno <- compute_ale(
model = mod_cox,
newdata = veteran,
feature = "karno",
times = c(100, 200, 300)
)
plot_ale(ale_karno, feature = "karno", which = "per_time")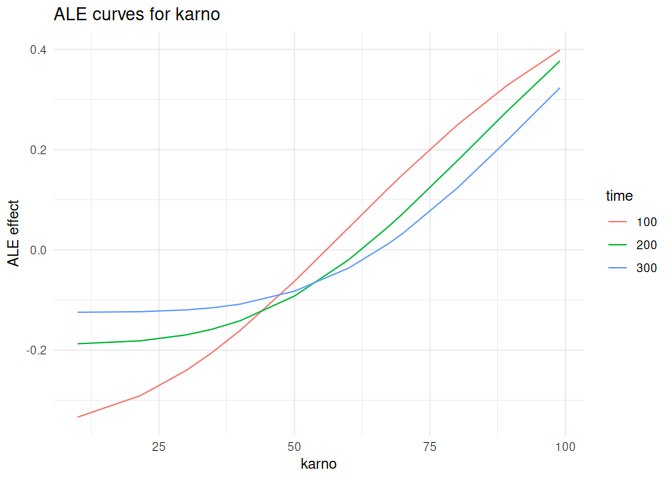
plot_ale(ale_karno, feature = "karno", which = "integrated", smooth = TRUE)
#> `geom_smooth()` using formula = 'y ~ x'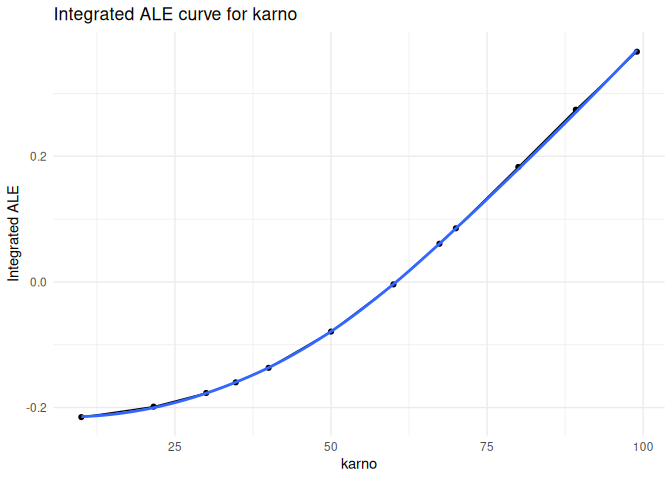
Local surrogate explanation
local_surrogate <- compute_surrogate(
model = mod_cox,
newdata = veteran[1, , drop = FALSE],
baseline_data = veteran,
times = c(100, 200, 300),
target_time = 200,
k = 5
)
local_surrogate
#> feature feature_value effect target_time
#> 1 karno 60 0.491034890 200
#> 2 celltype squamous 0.189632633 200
#> 3 age 69 0.120729843 200
#> 4 diagtime 7 0.001800378 200
#> 5 prior 0 0.000000000 200
plot_surrogate(local_surrogate, top_n = 10)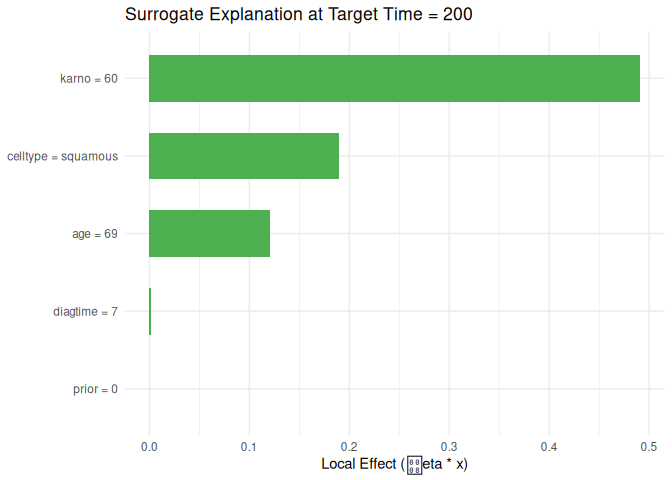
Tree surrogate
tree_surrogate <- compute_tree_surrogate(
model = mod_cox,
data = veteran,
times = c(100, 200, 300)
)
plot_tree_surrogate(tree_surrogate, type = "importance", top_n = 5)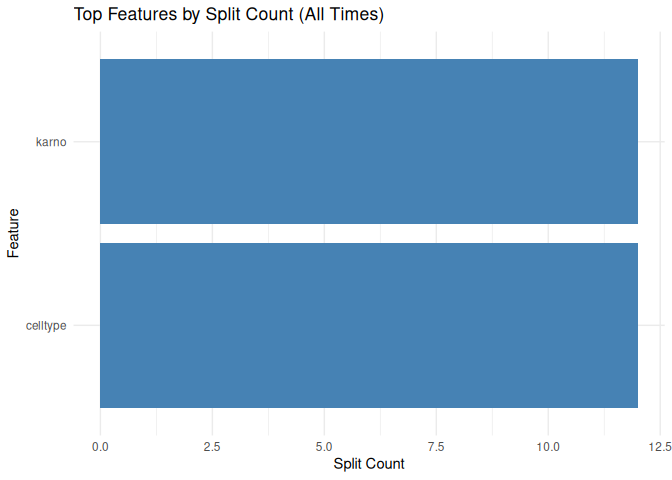
# plot_tree_surrogate(tree_surrogate, type = "tree")Permutation variable importance
varimp_res <- compute_varimp(
model = mod_cox,
times = c(100, 200, 300),
metric = "ibs",
n_repetitions = 5,
seed = 123
)
varimp_res
#> # A tibble: 6 × 5
#> feature importance importance_05 importance_95 scaled_importance
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 karno 0.0624 0.192 0.223 100
#> 2 celltype 0.0446 0.185 0.201 71.5
#> 3 age -0.00180 0.144 0.146 2.89
#> 4 trt 0 0.147 0.147 0
#> 5 diagtime 0 0.147 0.147 0
#> 6 prior 0 0.147 0.147 0
plot_varimp(varimp_res)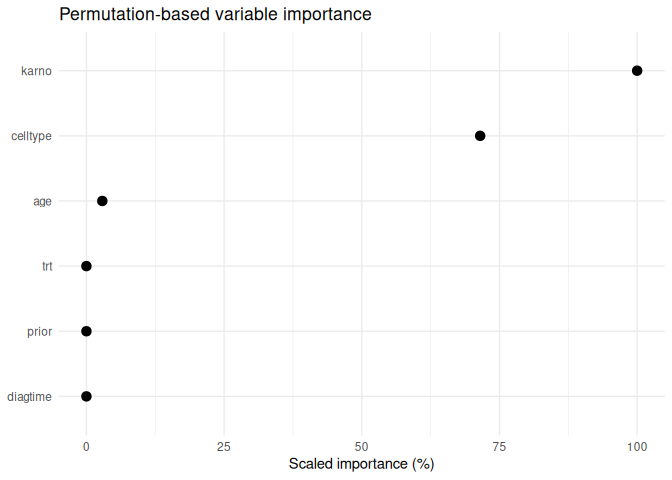
Feature interactions
interaction_1way <- compute_interactions(
model = mod_cox,
data = veteran,
times = c(100, 200, 300),
target_time = 200,
type = "1way"
)
interaction_heatmap <- compute_interactions(
model = mod_cox,
data = veteran,
times = c(100, 200, 300),
target_time = 200,
type = "heatmap"
)
interaction_time <- compute_interactions(
model = mod_cox,
data = veteran,
times = c(100, 200, 300),
type = "time"
)
plot_interactions(interaction_1way, type = "1way")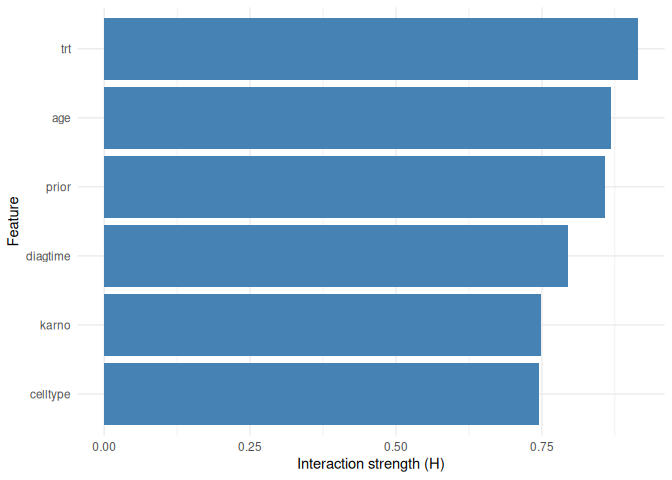
plot_interactions(interaction_heatmap, type = "heatmap")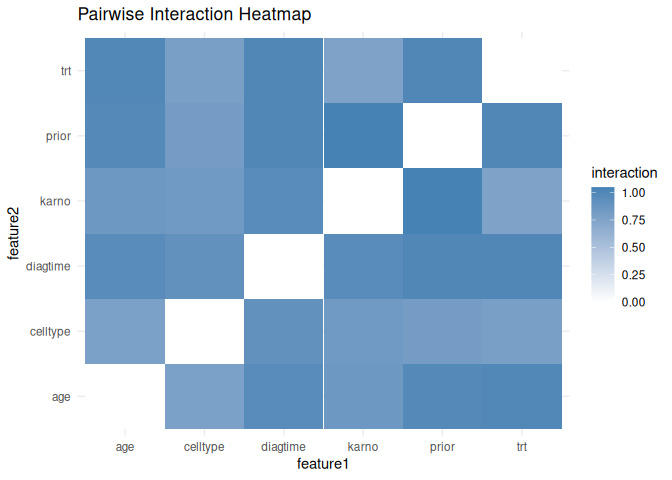
plot_interactions(interaction_time, type = "time")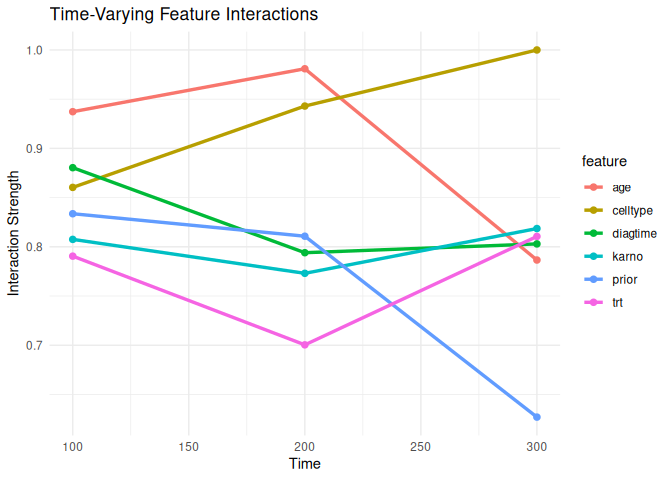
Counterfactual explanations
counterfactuals <- compute_counterfactual(
model = mod_cox,
newdata = veteran[1, , drop = FALSE],
times = c(100, 200, 300),
target_time = 200,
features_to_change = c("age", "karno", "diagtime"),
cost_penalty = 0.01
)
counterfactuals
#> feature original_value suggested_value survival_gain change_cost
#> 1 karno 60 81.0202 0.2347 21.0202
#> 2 diagtime 7 7.0808 0.0000 0.0808
#> 3 age 69 69.1313 0.0003 0.1313
#> penalized_gain
#> 1 0.0245
#> 2 -0.0008
#> 3 -0.00105. Calibration
compute_calibration(
model = mod_cox, data = veteran,
time = "time", status = "status",
eval_time = 80, n_bins = 10, n_boot = 30
) |> plot_calibration()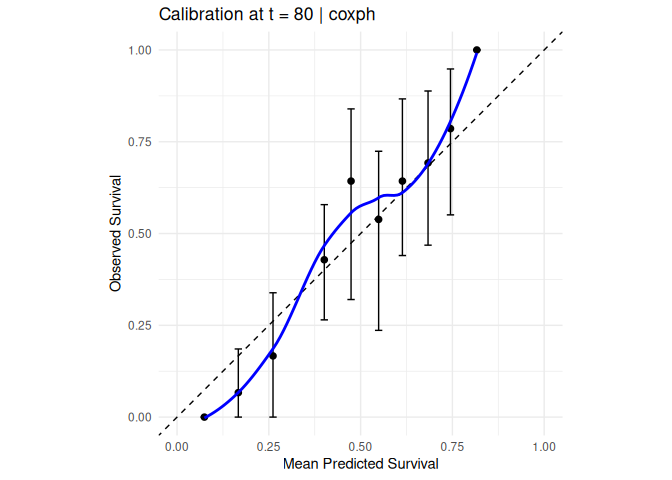
citation("survalis")These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.