The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
stringfish is a framework for string and sequence
operations using the ALTREP system (introduced in R 3.5) as a way to
represent R objects using custom memory layout.
This package has two primary goals:
stringfish currently provides two ALTREP backends with
the same semantics: sf_vec, a simple vector of string
objects, and slice_store, which stores strings within large
contiguous blocks of memory. They make different storage tradeoffs, but
the same stringfish operations work across both.
For text data, stringfish is intentionally UTF-8-centric
outside of explicit byte mode, so conversions, comparisons, and ALTREP
views stay consistent across normal R vectors and both backends.
install.packages("stringfish", type="source", configure.args="--with-simd=AVX2")The simplest way to show the utility of the ALTREP framework is
through a quick benchmark comparing stringfish and base
R.
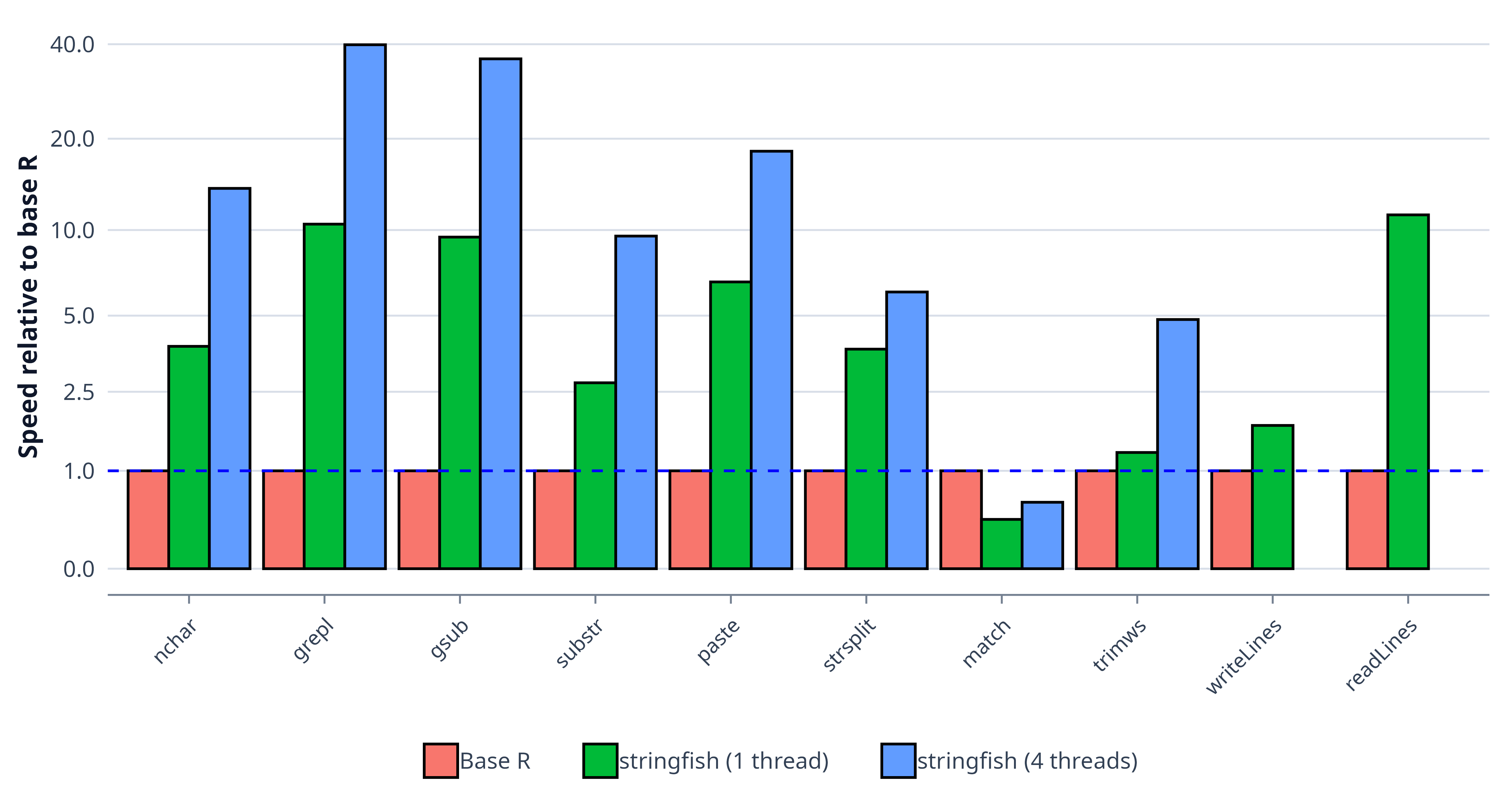
On favorable workloads, some functions in stringfish can
be more than an order of magnitude faster than vectorized base R
operations, and built-in multithreading can widen that gap further. On
large text datasets, this can turn minutes of computation into
seconds.
A list of implemented stringfish functions and analogous
base R functions:
sf_iconv (iconv)sf_nchar (nchar)sf_substr (substr)sf_paste (paste0)sf_collapse (paste0)sf_readLines (readLines)sf_writeLines (writeLines)sf_grepl (grepl)sf_gsub (gsub)sf_toupper (toupper)sf_tolower (tolower)sf_starts (startsWith)sf_ends (endsWith)sf_trim (trimws)sf_split (strsplit)sf_match (match for strings only)sf_compare/sf_equals (==,
ALTREP-aware semantic string equality)sf_concat/sfc (c)Utility functions:
sf_vector_create – creates a new empty
sf_vec-backed stringfish vectorsf_vector – backwards-compatible alias for
sf_vector_createslice_store_create – creates a new empty
slice_store-backed stringfish vectorslice_store_create_with_size – creates a
slice_store-backed stringfish vector with an explicit
initial slice sizesf_assign – assign strings into a
stringfish vector in place (like
x[i] <- "mystring")convert_to_sf_vector – converts a character vector to a
stringfish vectorconvert_to_slice_store – converts a character vector to
a stringfish slice storeget_string_type – determines string type (whether
ALTREP or normal)materialize – converts any ALTREP object into a normal
R objectrandom_strings – creates random strings as either a
stringfish or normal R vectorstring_identical – compares strings either semantically
or exactly across encodingsIn addition, many R operations in base R and other packages are already ALTREP-aware (i.e. they don’t cause materialization). Functions that subset or index into string vectors generally do not materialize.
sampleheadtail[ – e.g. x[20:30]stringfish functions are not intended to exactly
replicate their base R analogues. One difference is that
subject parameters are always the first argument, which is
easier to use with pipes. E.g.,
gsub(pattern, replacement, subject) becomes
sf_gsub(subject, pattern, replacement).
stringfish as a framework is intended to be easily
extensible. Stringfish vectors can be worked into Rcpp
scripts or even into other packages. The example below creates an
sf_vec-backed output because it is simple and direct, but
the same indexing semantics work across both backends.
Below is a detailed Rcpp script that creates a function
to alternate upper and lower case of strings.
// [[Rcpp::depends(stringfish)]]
#include <Rcpp.h>
#include "sf_external.h"
using namespace Rcpp;
// [[Rcpp::export]]
SEXP sf_alternate_case(SEXP x) {
// Iterate through a character vector using the RStringIndexer class
// If the input vector x is a stringfish character vector it will do so without materialization
RStringIndexer r(x);
size_t len = r.size();
// Create an output stringfish vector
// Like all R objects, it must be protected from garbage collection
SEXP output = PROTECT(sf_vector_create(len));
// Obtain a reference to the underlying output data
sf_vec_data & output_data = sf_vec_data_ref(output);
// You can use range based for loop via an iterator class that returns RStringIndexer::rstring_info e
// rstring info is a struct containing const char * ptr, int len, and an encoding flag
// ptr should be treated as a byte pointer plus length, not as a null-terminated C string
// a NA string is represented by a nullptr
// Alternatively, access the data via the function r.getCharLenCE(i)
size_t i = 0;
for(auto e : r) {
// check if string is NA and go to next if it is
if(e.ptr == nullptr) {
i++; // increment output index
continue;
}
// Create a temporary output string and process the results.
// This example intentionally toggles ASCII letters only.
std::string temp(e.len, '\0');
bool case_switch = false;
for(int j=0; j<e.len; j++) {
if((e.ptr[j] >= 65) && (e.ptr[j] <= 90)) { // char j is upper case
if((case_switch = !case_switch)) { // check if we should convert to lower case
temp[j] = e.ptr[j] + 32;
continue;
}
} else if((e.ptr[j] >= 97) && (e.ptr[j] <= 122)) { // char j is lower case
if(!(case_switch = !case_switch)) { // check if we should convert to upper case
temp[j] = e.ptr[j] - 32;
continue;
}
} else if(e.ptr[j] == 32) {
case_switch = false;
}
temp[j] = e.ptr[j];
}
// Create a new vector element sfstring and insert the processed string into the stringfish vector
// sfstring has three constructors, 1) taking a std::string and encoding,
// 2) a char pointer and encoding, or 3) a CHARSXP object (e.g. sfstring(NA_STRING))
output_data[i] = sfstring(temp, e.enc);
i++; // increment output index
}
// Finally, call unprotect and return result
UNPROTECT(1);
return output;
}Example function call:
sf_alternate_case("hello world")
[1] "hElLo wOrLd"These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.