The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
splitGraph is an R package for representing biomedical
dataset structure as a typed dependency graph so that leakage-relevant
relationships can be made explicit, validated, queried, and converted
into deterministic split constraints.
It does not fit models, run preprocessing pipelines, or generate resamples by itself. Its job is to encode dataset structure before evaluation so that overlap, provenance, and time-ordering assumptions are inspectable instead of implicit.

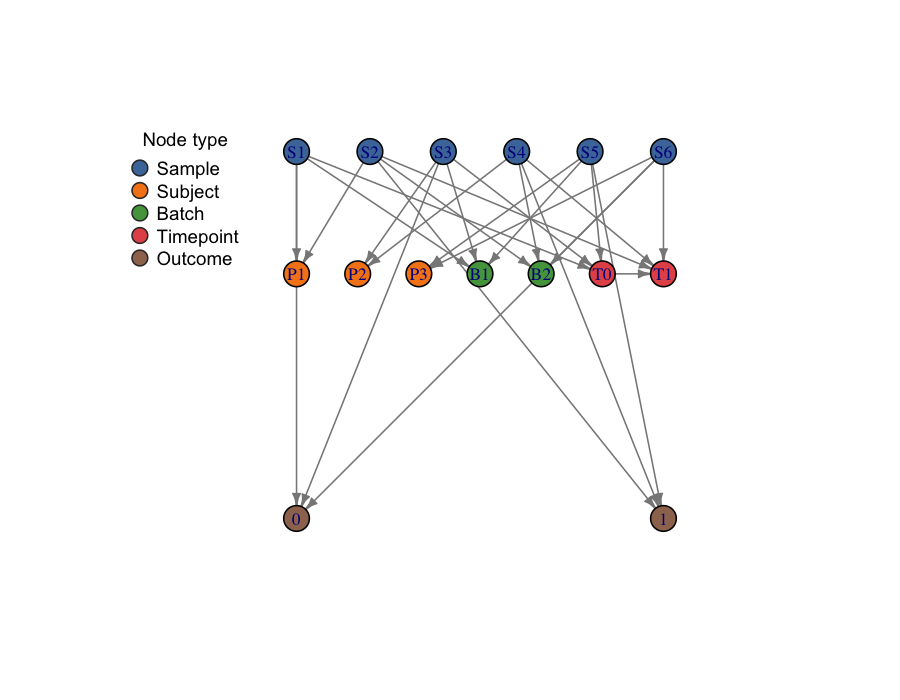
The plot above shows six samples (blue) that share three subjects
(orange), two batches (green), two timepoints (red), and two outcome
classes (brown). A plain vfold_cv on this dataset would
violate subject, batch, and time structure at the same time —
and that is exactly what the graph is designed to make visible.
In biomedical evaluation workflows, leakage often comes from dataset structure rather than obvious coding mistakes. Samples may share:
If those relationships are not modeled explicitly, a train/test split can look correct while still violating the intended scientific separation.
splitGraph makes those dependencies first-class
objects.
Does:
graph_from_metadata()igraphvalidation_overrides mechanism for explicit
exceptionsquery_paths())split_specdependency_graph and
split_spec so handoff objects are portable across sessions
and languages (write_*() / read_*(), requires
jsonlite)plot() method with per-type colors and a
node-type legendprint(), summary(), and
as.data.frame() on all core S3 objectsDoes not:
rsample does that)The package is intentionally narrow: dataset dependency structure for leakage-aware evaluation design.
From GitHub:
install.packages("remotes")
remotes::install_github("selcukorkmaz/splitGraph")To use the JSON serialization API, also install
jsonlite:
install.packages("jsonlite")The fastest path is graph_from_metadata(), which
auto-detects canonical columns in a metadata frame and assembles a
validated dependency_graph:
library(splitGraph)
meta <- data.frame(
sample_id = c("S1", "S2", "S3", "S4", "S5", "S6"),
subject_id = c("P1", "P1", "P2", "P2", "P3", "P3"),
batch_id = c("B1", "B2", "B1", "B2", "B1", "B2"),
timepoint_id = c("T0", "T1", "T0", "T1", "T0", "T1"),
time_index = c(0, 1, 0, 1, 0, 1),
outcome_id = c("ctrl", "case", "ctrl", "case", "case", "ctrl")
)
g <- graph_from_metadata(meta, graph_name = "demo")
plot(g)
validation <- validate_graph(g)
subject_constraint <- derive_split_constraints(g, mode = "subject")
spec <- as_split_spec(subject_constraint, graph = g)
validate_split_spec(spec)
summarize_leakage_risks(g, constraint = subject_constraint, split_spec = spec)
# Persist the spec for a downstream consumer (R or non-R):
path <- tempfile(fileext = ".json")
write_split_spec(spec, path)
spec2 <- read_split_spec(path)For full control over node labels, attribute columns, and the
feature-set provenance edges, use create_nodes() /
create_edges() / build_dependency_graph()
directly. graph_from_metadata() auto-builds the six
sample-rooted canonical edges, timepoint_precedes, and the
appropriate outcome edge (sample_has_outcome by default, or
subject_has_outcome when
outcome_scope = "subject"). The
featureset_generated_from_study and
featureset_generated_from_batch edges always require the
explicit constructor path.
split_spec is the tool-agnostic handoff object produced
by as_split_spec(). splitGraph does not know
about any particular resampling package — downstream consumers are
expected to provide their own adapters so that splitGraph
stays neutral and has no runtime dependency on them.
The typical end-to-end flow is:
graph_from_metadata(meta) → typed
dependency_graphderive_split_constraints(g, mode = ...) →
split_constraintas_split_spec(constraint, graph = g) →
split_specwrite_split_spec(spec, path) → JSON, for
cross-session or cross-language handoffThe sample_data frame carried by split_spec
exposes exactly what an adapter needs: sample_id for
joining against the observation frame, group_id for grouped
resampling, batch_group / study_group for
blocking, and order_rank for ordered evaluation. An adapter
can be built on top of, for example,
rsample::group_vfold_cv() (grouped CV keyed to
group_id) or rsample::rolling_origin()
(ordered evaluation keyed to order_rank).
For three small, self-contained adapter examples (a base-R LOGO
adapter, plus illustrative rsample::group_vfold_cv() and
rsample::rolling_origin() adapters), see the
Adapter cookbook vignette:
vignette("adapter-cookbook", package = "splitGraph")Sample, Subject, Batch,
Study, Timepoint, Assay,
FeatureSet, Outcomesample_belongs_to_subjectsample_processed_in_batchsample_from_studysample_collected_at_timepointsample_measured_by_assaysample_uses_featuresetsample_has_outcomesubject_has_outcometimepoint_precedesfeatureset_generated_from_studyfeatureset_generated_from_batchgraph_node_set, graph_edge_set,
dependency_graph, depgraph_validation_report,
graph_query_result, split_constraint,
split_spec, split_spec_validation,
leakage_risk_summary.
| Layer | Functions |
|---|---|
| Ingestion and construction | ingest_metadata(), graph_from_metadata(),
create_nodes(), create_edges(),
build_dependency_graph(), dependency_graph(),
as_igraph() |
| Validation | validate_graph() (with
validation_overrides),
validate_split_spec() |
| Queries | query_node_type(), query_edge_type(),
query_neighbors(), query_paths() (capped by
default), query_shortest_paths(),
detect_dependency_components(),
detect_shared_dependencies() |
| Constraint derivation | derive_split_constraints(),
grouping_vector() |
| Split-spec translation | as_split_spec(),
summarize_leakage_risks() |
| Serialization (JSON) | write_dependency_graph(),
read_dependency_graph(), write_split_spec(),
read_split_spec() |
query_node_type(g, "Subject")
query_edge_type(g, "sample_processed_in_batch")
query_neighbors(g, node_ids = "sample:S1", edge_types = "sample_belongs_to_subject")
detect_shared_dependencies(g, via = "Batch")
detect_dependency_components(g, via = c("Subject", "Batch"))subject_constraint <- derive_split_constraints(g, mode = "subject")
batch_constraint <- derive_split_constraints(g, mode = "batch")
study_constraint <- derive_split_constraints(g, mode = "study")
time_constraint <- derive_split_constraints(g, mode = "time")
strict_composite <- derive_split_constraints(
g, mode = "composite", strategy = "strict",
via = c("Subject", "Batch")
)
rule_based_composite <- derive_split_constraints(
g, mode = "composite", strategy = "rule_based",
priority = c("batch", "study", "subject", "time")
)Both core handoff objects can be written to a stable,
schema-versioned JSON format and read back, so a
dependency_graph or split_spec is portable
across R sessions and across language boundaries.
graph_path <- tempfile(fileext = ".json")
spec_path <- tempfile(fileext = ".json")
write_dependency_graph(g, graph_path)
write_split_spec(spec, spec_path)
g2 <- read_dependency_graph(graph_path)
spec2 <- read_split_spec(spec_path)The JSON schema is documented under
?write_dependency_graph and ?write_split_spec.
Each file carries a schema_version field; reading a file
written under a different schema version emits a warning but still
loads. NA values in sample_data round-trip as
JSON null. The jsonlite package (a
Suggests dep) must be installed.
plot(g) renders a typed, layered layout with per-type
node colors and an auto-generated node-type legend. Layers: Sample
(top), peer dependencies (Subject / Batch / Study / Timepoint) in the
middle band, Assay / FeatureSet next, Outcome (bottom).
plot(g) # typed layered layout (default)
plot(g, layout = "sugiyama") # alternative hierarchical layout
plot(g, show_labels = FALSE) # hide node labels on dense graphs
plot(g, legend = FALSE) # suppress the legend
plot(g, legend_position = "bottomright")
plot(g, node_colors = c(Sample = "#000000")) # override type colorscitation("splitGraph")produces:
Korkmaz S (2026). splitGraph: Dataset Dependency Graphs for Leakage-Aware Evaluation. R package version 0.2.0. https://github.com/selcukorkmaz/splitGraph
MIT. See LICENSE.
The package prefers explicit failure over silent guessing. In particular:
validation_overrides
(e.g. allow_multi_subject_samples); the same override is
honored by both validate_graph() and
derive_split_constraints(mode = "subject")query_paths() defaults to a finite path-length cap so
traversal cannot explode on dense graphs; pass
max_length = Inf to opt outschema_version warns rather than failing
silentlyThese binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.