The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
This vignette introduces the use of the refundBayes
package with simple examples for: (1) Bayesian Scalar-on-Function
Regression (SoFR), (2) Bayesian Functional Cox Rergession (FCox), and
(3) Bayesian Function-on-Scalar Rergession (FoSR). The
refundBayes package provides users with a convenient way to
perform Bayesian functional data analysis using Stan, a state-of-the-art
Bayesian analysis language. The syntax of refundBayes is
designed to resemble that of mgcv::gam, but it employs
Bayesian posterior inference and includes additional arguments specific
to the Stan implementation.
library(remotes)
remotes::install_github("https://github.com/ZirenJiang/refundBayes")## Skipping install of 'refundBayes' from a github remote, the SHA1 (5a1f14ca) has not changed since last install.
## Use `force = TRUE` to force installation## Read the example data
data.SoFR = readRDS("data/example_data_sofr.rds")
## Run Bayesian SOFR with the sofr_bayes() function
refundBayes_SoFR = refundBayes::sofr_bayes(y ~ X1+s(tmat, by=lmat*wmat, bs="cc", k=10),
data = data.SoFR,
family = binomial(),
runStan = TRUE, # Whether automatically run Stan program.
niter = 1500, # Total number of posterior sampling.
nwarmup = 500, # Burn-in value.
nchain = 3, # Number of parallel computed chains for posterior sampling.
ncores = 3
)
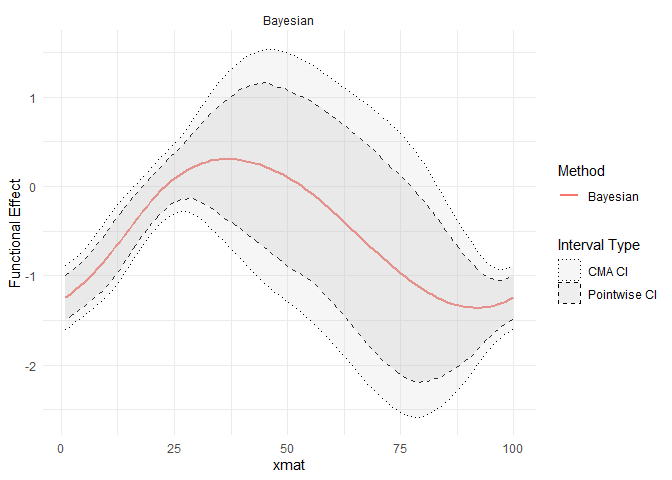
## Visualization of the fitted functional coefficient
library(ggplot2)
plot(refundBayes_SoFR)## Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
## ℹ Please use `linewidth` instead.
## ℹ The deprecated feature was likely used in the refundBayes package.
## Please report the issue to the authors.
## This warning is displayed once every 8 hours.
## Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
## generated.
## [[1]]
mean.curve.est = apply(refundBayes_SoFR$func_coef[[1]], 2, mean)
upper.curve.est = apply(refundBayes_SoFR$func_coef[[1]], 2, function(x){quantile(x, prob = 0.975)})
lower.curve.est = apply(refundBayes_SoFR$func_coef[[1]], 2, function(x){quantile(x, prob = 0.025)})
library(mgcv)
## This is mgcv 1.9-1. For overview type 'help("mgcv-package")'.fit_freq1 = gam(y ~ s(tmat, by=lmat*wmat, bs="cc", k=10)+X1, data=data.SoFR, family="binomial")
fit_freq2 = gam(y ~ s(tmat, by=lmat*wmat, bs="cc", k=10)+X1, data=data.SoFR, family="binomial", method = "REML")
plotfot=plot(fit_freq1)
plotfot2=plot(fit_freq2,unconditional = TRUE)
T_num = 100
plotdata=data.frame(value=c(mean.curve.est,
upper.curve.est,
lower.curve.est,
plotfot[[1]]$fit,
plotfot[[1]]$fit+plotfot[[1]]$se,
plotfot[[1]]$fit-plotfot[[1]]$se,
plotfot2[[1]]$fit,
plotfot2[[1]]$fit+plotfot2[[1]]$se,
plotfot2[[1]]$fit-plotfot2[[1]]$se),
xmat=c(rep(1: T_num,3),
rep(plotfot[[1]]$x*100,6)),
Method=c(rep("Bayesian",T_num),
rep("Bayesian",T_num),
rep("Bayesian",T_num),
rep("Frequentist - GCV",100),
rep("Frequentist - GCV",100),
rep("Frequentist - GCV",100),
rep("Frequentist - REML",100),
rep("Frequentist - REML",100),
rep("Frequentist - REML",100)),
type=c(rep("Estimate",T_num),
rep("CI_upper",T_num),
rep("CI_lower",T_num),
rep("Estimate",100),
rep("CI_upper",100),
rep("CI_lower",100),
rep("Estimate",100),
rep("CI_upper",100),
rep("CI_lower",100)))
library(ggplot2)
ggplot(plotdata,aes(y=value,x=xmat))+geom_line(aes(linetype=type,color=Method))+
ylab("Functional effect")+xlab("Time index")+
#scale_x_continuous(breaks=seq(0,1,by=3))+
scale_linetype_manual(values=c("twodash", "longdash","solid"),name="Line Type")+
theme_minimal()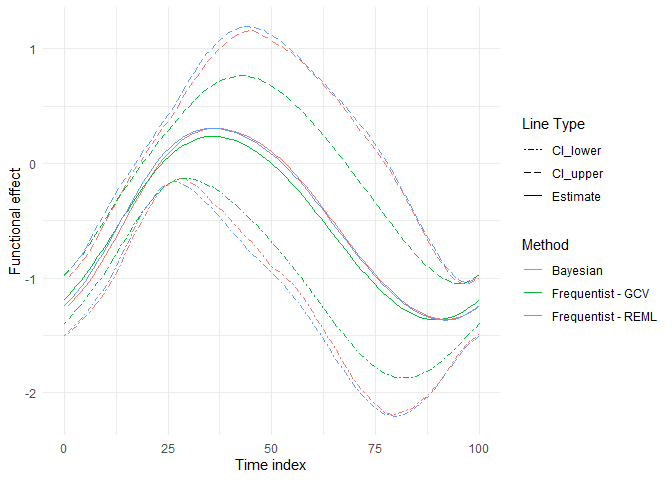
## Read the example data
Func_Cox_Data = readRDS("data/example_data_Cox.rds")
Func_Cox_Data$wmat = Func_Cox_Data$MIMS
Func_Cox_Data$cens = 1 - Func_Cox_Data$event
## Run Bayesian SOFR with the sofr_bayes() function
refundBayes_FCox = refundBayes::fcox_bayes(survtime ~ X1+s(tmat, by=lmat*wmat, bs="cc", k=10),
data = Func_Cox_Data,
cens = Func_Cox_Data$cens,
runStan = TRUE, # Whether automatically run Stan program.
niter = 1500, # Total number of posterior sampling.
nwarmup = 500, # Burn-in value.
nchain = 3, # Number of parallel computed chains for posterior sampling.
ncores = 3
)## Warning: There were 6 divergent transitions after warmup. See
## https://mc-stan.org/misc/warnings.html#divergent-transitions-after-warmup
## to find out why this is a problem and how to eliminate them.
## Warning: Examine the pairs() plot to diagnose sampling problems## Visualization of the fitted functional coefficient
library(ggplot2)
plot(refundBayes_FCox)## [[1]]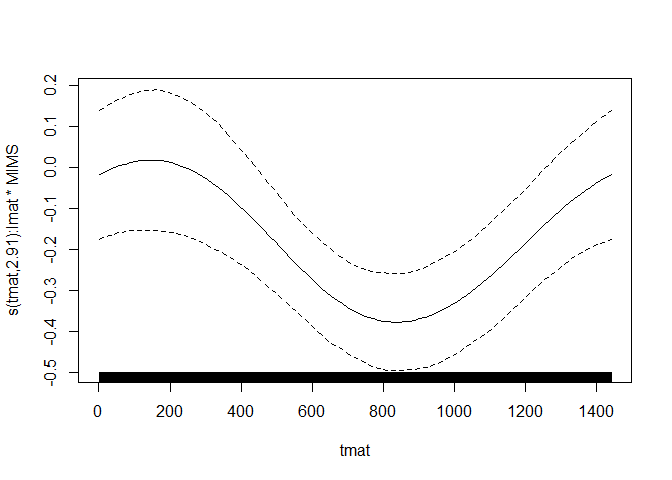
## Read the example data
FoSR_exp_data = readRDS("data/example_data_FoSR.rds")
FoSR_exp_data$y = FoSR_exp_data$MIMS
## Run Bayesian SOFR with the sofr_bayes() function
refundBayes_FoSR = refundBayes::fosr_bayes(y ~ X,
data = FoSR_exp_data,
runStan = TRUE, # Whether automatically run Stan program.
niter = 1500, # Total number of posterior sampling.
nwarmup = 500, # Burn-in value.
nchain = 1, # Number of parallel computed chains for posterior sampling.
ncores = 1
)##
## SAMPLING FOR MODEL 'anon_model' NOW (CHAIN 1).
## Chain 1:
## Chain 1: Gradient evaluation took 0.001614 seconds
## Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 16.14 seconds.
## Chain 1: Adjust your expectations accordingly!
## Chain 1:
## Chain 1:
## Chain 1: Iteration: 1 / 1500 [ 0%] (Warmup)
## Chain 1: Iteration: 150 / 1500 [ 10%] (Warmup)
## Chain 1: Iteration: 300 / 1500 [ 20%] (Warmup)
## Chain 1: Iteration: 450 / 1500 [ 30%] (Warmup)
## Chain 1: Iteration: 501 / 1500 [ 33%] (Sampling)
## Chain 1: Iteration: 650 / 1500 [ 43%] (Sampling)
## Chain 1: Iteration: 800 / 1500 [ 53%] (Sampling)
## Chain 1: Iteration: 950 / 1500 [ 63%] (Sampling)
## Chain 1: Iteration: 1100 / 1500 [ 73%] (Sampling)
## Chain 1: Iteration: 1250 / 1500 [ 83%] (Sampling)
## Chain 1: Iteration: 1400 / 1500 [ 93%] (Sampling)
## Chain 1: Iteration: 1500 / 1500 [100%] (Sampling)
## Chain 1:
## Chain 1: Elapsed Time: 51.492 seconds (Warm-up)
## Chain 1: 9.381 seconds (Sampling)
## Chain 1: 60.873 seconds (Total)
## Chain 1:
## Warning: The largest R-hat is NA, indicating chains have not mixed.
## Running the chains for more iterations may help. See
## https://mc-stan.org/misc/warnings.html#r-hat
## Warning: Bulk Effective Samples Size (ESS) is too low, indicating posterior means and medians may be unreliable.
## Running the chains for more iterations may help. See
## https://mc-stan.org/misc/warnings.html#bulk-ess
## Warning: Tail Effective Samples Size (ESS) is too low, indicating posterior variances and tail quantiles may be unreliable.
## Running the chains for more iterations may help. See
## https://mc-stan.org/misc/warnings.html#tail-ess## Visualization of the fitted functional coefficient
library(ggplot2)
plot(refundBayes_FoSR)## Warning in ggplot2::geom_line(aes(type = .data$type, color = .data$type)):
## Ignoring unknown aesthetics: type
## [[1]]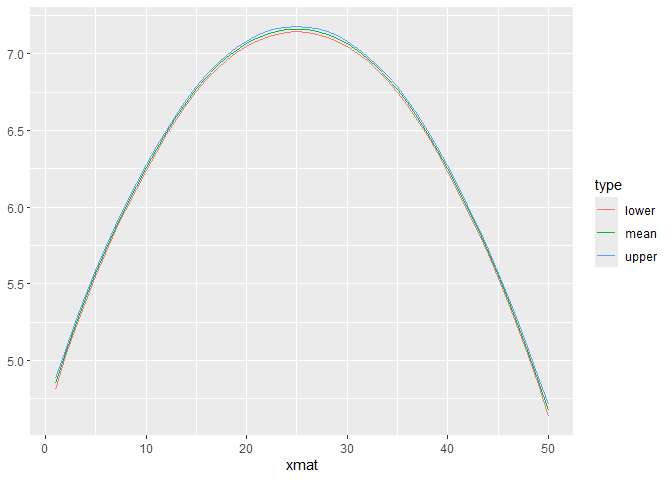
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.