The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
Explore various dependencies of a package(s) (on the Comprehensive R Archive Network Like repositories).
Start with init() which gets metadata from a CRAN like
repository.
The package offers four functions:
get_dependencies(): Get direct or indirect dependencies
of a set of packages at desired depth (as a dataframe).get_neighborhood(): Get both direct and indirect of a
set of packages at desired depth (as a dataframe).as_graph(): Get metadata embedded igraph
equivalent of a dependency dataframe.plot(): Static plot of dependency graph.suppressPackageStartupMessages(library("dplyr"))
#> Warning: package 'dplyr' was built under R version 4.4.3
init()
#> ℹ Fetching package metadata from repositories ...
#> ℹ Computing package dependencies ...
#> ✔ Done!
# what does `mboost` package do
packmeta |> filter(Package == "mboost") |> pull(Description) |> cat()
#> Functional gradient descent algorithm
#> (boosting) for optimizing general risk functions utilizing
#> component-wise (penalised) least squares estimates or regression
#> trees as base-learners for fitting generalized linear, additive
#> and interaction models to potentially high-dimensional data.
#> Models and algorithms are described in <doi:10.1214/07-STS242>,
#> a hands-on tutorial is available from <doi:10.1007/s00180-012-0382-5>.
#> The package allows user-specified loss functions and base-learners.
# Get direct 'import' dependencies of `mboost` two levels deeper
get_dependencies("mboost", level = 2, relation = "Imports")
#> # A tibble: 58 × 3
#> pkg_1 relation pkg_2
#> <chr> <fct> <chr>
#> 1 BayesX Imports coda
#> 2 BayesX Imports colorspace
#> 3 BayesX Imports interp
#> 4 BayesX Imports sf
#> 5 BayesX Imports sp
#> 6 BayesX Imports splines
#> 7 Matrix Imports grid
#> 8 Matrix Imports grid
#> 9 Matrix Imports grid
#> 10 Matrix Imports lattice
#> # ℹ 48 more rows
# Get neighborhood (direct and indirect) dependencies (of all types) of `mboost` one level deeper
get_neighborhood("mboost", level = 1)
#> # A tibble: 222 × 3
#> pkg_1 relation pkg_2
#> <chr> <fct> <chr>
#> 1 FDboost Depends mboost
#> 2 InvariantCausalPrediction Depends mboost
#> 3 TH.data Depends MASS
#> 4 TH.data Depends survival
#> 5 boostrq Depends mboost
#> 6 boostrq Depends parallel
#> 7 boostrq Depends stabs
#> 8 catdata Depends MASS
#> 9 censored Depends survival
#> 10 expectreg Depends BayesX
#> # ℹ 212 more rows
# convert a dependency dataframe to a graph
get_neighborhood("mboost", level = 1) |> as_graph()
#> IGRAPH 146a4f2 DN-- 56 222 --
#> + attr: name (v/c), title (v/c), relation (e/c)
#> + edges from 146a4f2 (vertex names):
#> [1] FDboost ->mboost
#> [2] InvariantCausalPrediction->mboost
#> [3] TH.data ->MASS
#> [4] TH.data ->survival
#> [5] boostrq ->mboost
#> [6] boostrq ->parallel
#> [7] boostrq ->stabs
#> [8] catdata ->MASS
#> + ... omitted several edges
# plot a dependency graph
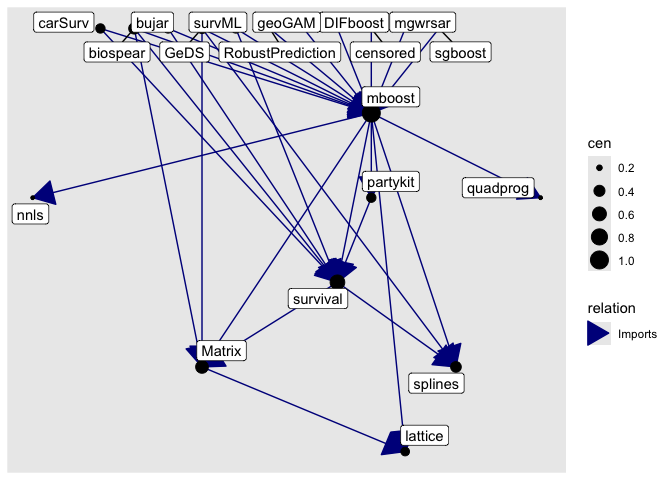
get_neighborhood("mboost", level = 1, relation = "Imports") |> as_graph() |> plot()
From CRAN:
pak::pkg_install("pkggraph")You can install the development version of pkggraph from GitHub with:
# install.packages("pak")
pak::pak("talegari/pkggraph")These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.