
The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
Audit trails track what happens at every step of a
dplyr pipeline by recording metadata-only snapshots: row counts, column
changes, NA totals, numeric shifts, and custom functions. Build a trail
by dropping taps into your pipe — transparent
pass-throughs that record a snapshot and let the data flow on unchanged.
Operation-aware taps, such as left_join_tap() and
filter_tap() go further, capturing match rates and drop
counts. The result is a structured trail you can print, diff, export as
HTML, or serialize to JSON. You can learn more in
vignette("tidyaudit").
tidyaudit also includes a diagnostic toolkit for
interactive data exploration — join validation, key checks, table
comparison, and more — described in
vignette("diagnostics").
library(tidyaudit)
library(dplyr)
set.seed(123)
orders <- data.frame(id = 1:100, amount = runif(100, 10, 500), region_id = sample(1:5, 100, TRUE))
regions <- data.frame(region_id = 1:4, name = c("North", "South", "East", "West"))
trail <- audit_trail("order_pipeline")
result <- orders |>
audit_tap(trail, "raw") |>
left_join_tap(regions, by = "region_id", .trail = trail, .label = "with_region") |>
filter_tap(amount > 100, .trail = trail, .label = "high_value", .stat = amount)
#> i filter_tap: amount > 100
#> Dropped 18 of 100 rows (18.0%)
#> Stat amount: dropped 1,062.191 of 25,429.39
print(trail)
#> -- Audit Trail: "order_pipeline" -----------------------------------------------
#> Created: 2026-02-21 14:36:35
#> Snapshots: 3
#>
#> # Label Rows Cols NAs Type
#> - ----------- ---- ---- --- ------------------------------------
#> 1 raw 100 3 0 tap
#> 2 with_region 100 4 23 left_join (many-to-one, 77% matched)
#> 3 high_value 82 4 20 filter (dropped 18 rows, 18%)
#>
#> Changes:
#> raw -> with_region: = rows, +1 cols, +23 NAs
#> with_region -> high_value: -18 rows, = cols, -3 NAs
audit_diff(trail, "raw", "high_value")
#> -- Audit Diff: "raw" -> "high_value" --
#>
#> Metric Before After Delta
#> ------ ------ ----- -----
#> Rows 100 82 -18
#> Cols 3 4 +1
#> NAs 0 20 +20
#>
#> Columns added: name
#>
#> Numeric shifts (common columns):
#> Column Mean before Mean after Shift
#> --------- ----------- ---------- ------
#> id 50.50 49.66 -0.84
#> amount 254.29 297.16 +42.87
#> region_id 3.08 3.05 -0.03Three taps. Three snapshots. A complete record of what the pipeline did to your data — and what it cost.
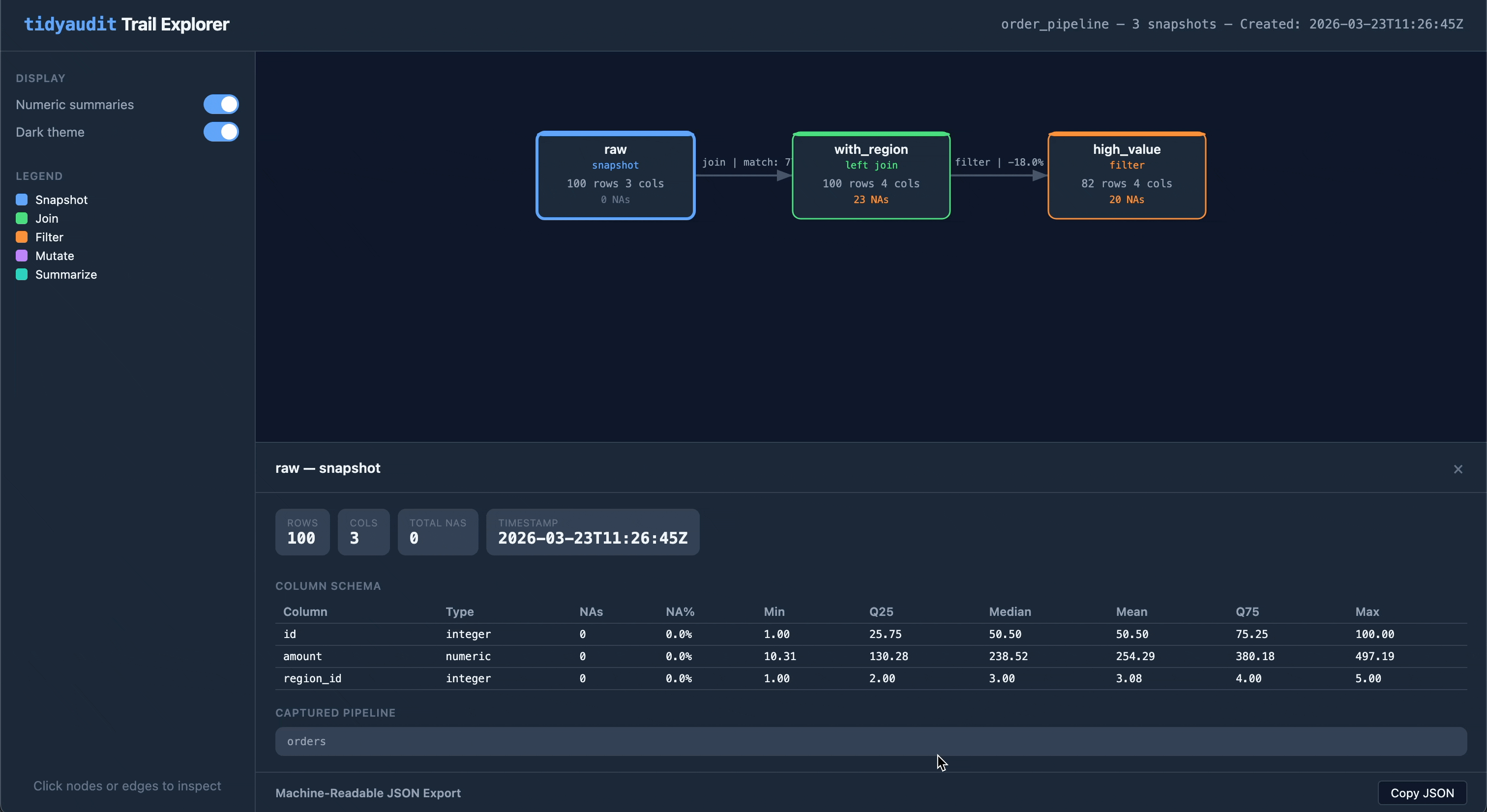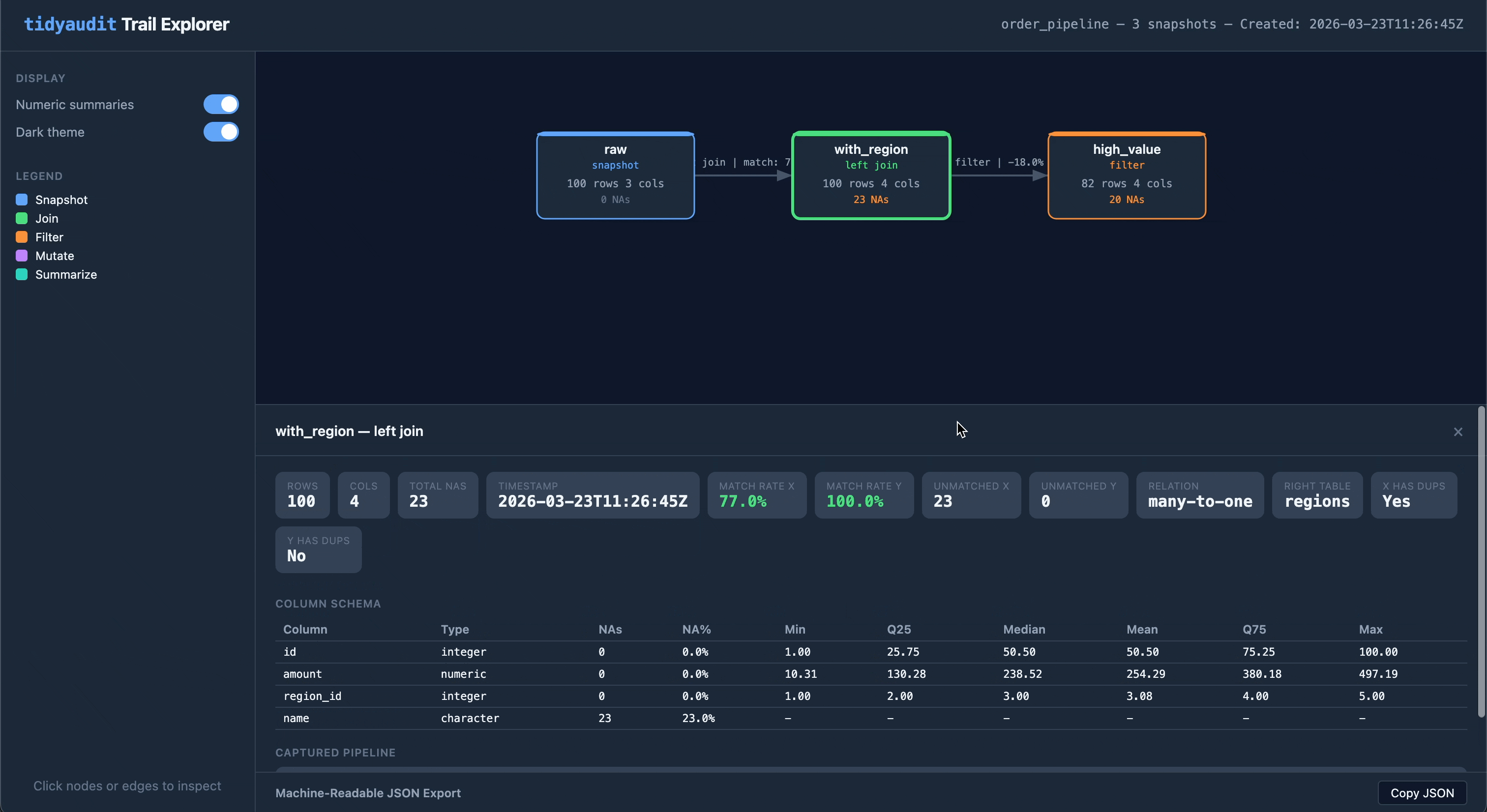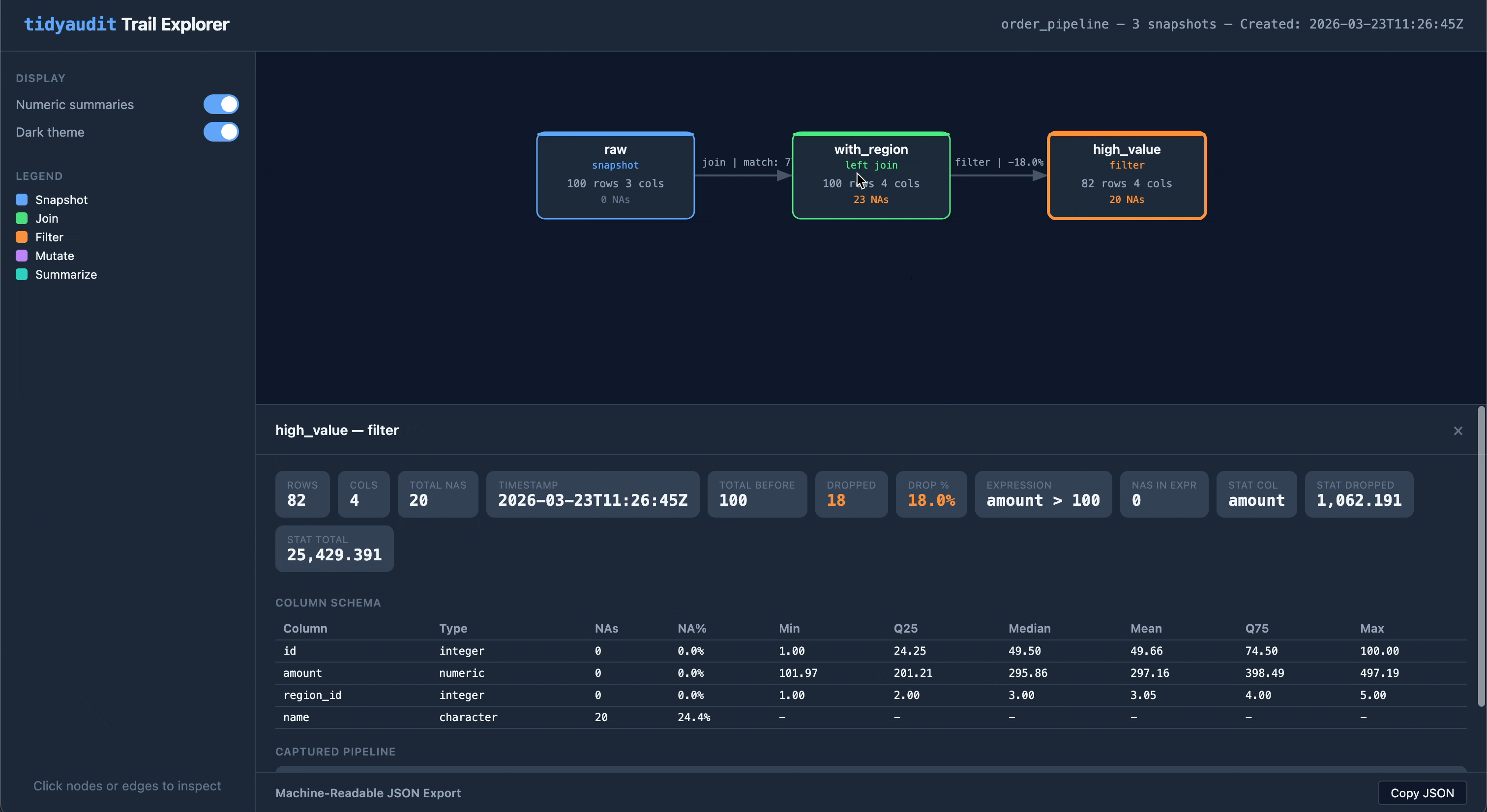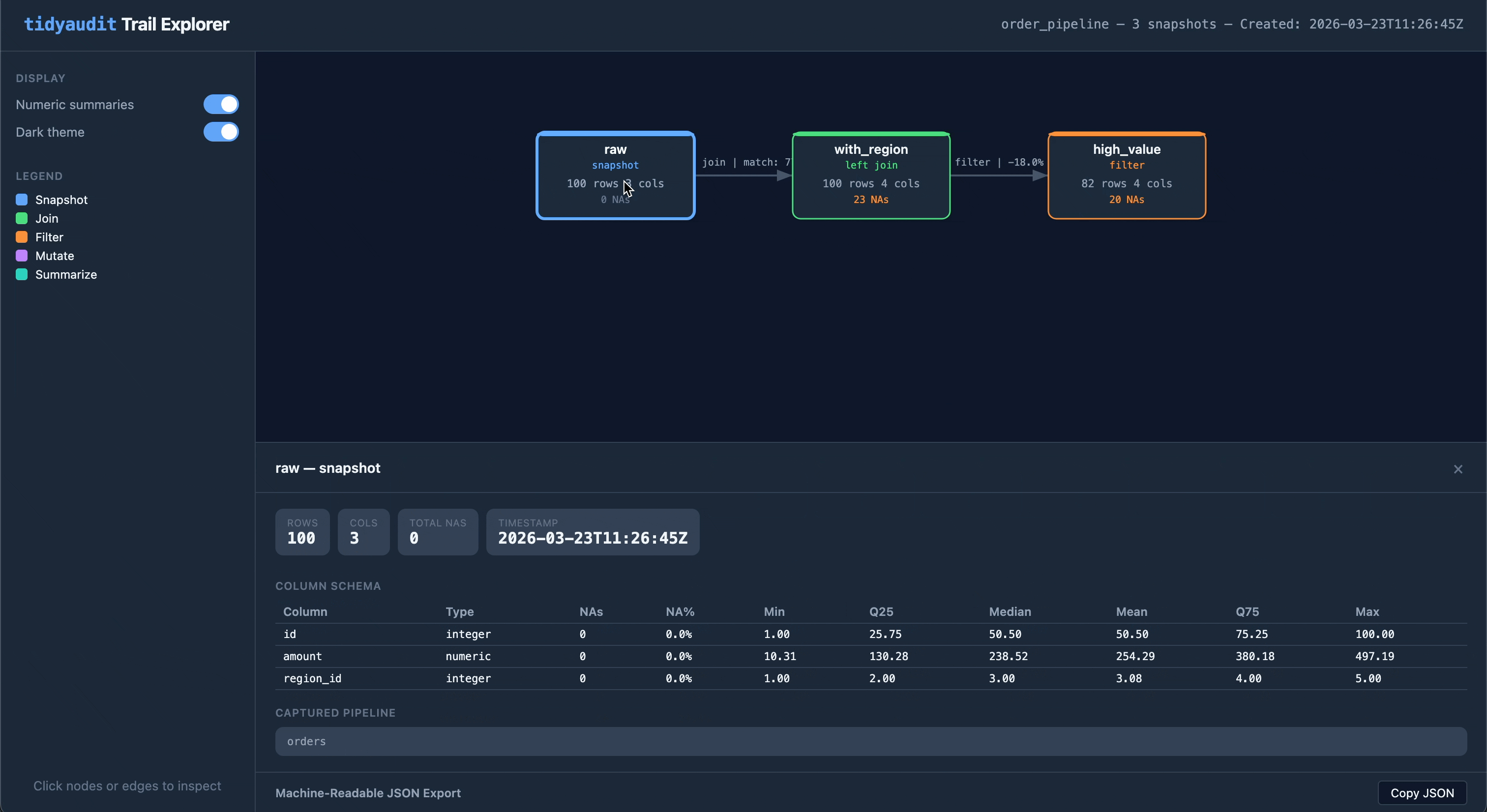
Share a trail as a self-contained HTML file — one file you can email, attach to a report, or drop into a compliance folder:
audit_export(trail, "order_pipeline.html")The output is an interactive flow diagram with clickable nodes and edges, light/dark theme toggle, and embedded JSON export. No server or internet required.
Build a structured timeline of your pipeline’s behavior. Drop taps into any dplyr pipe and get a traceable, diffable, exportable record of every step.
audit_trail() / audit_tap() — create a
trail and record snapshots inside pipesleft_join_tap(), filter_tap(), and friends
— operation-aware taps that capture match rates, drop
counts, stat impact, and relationship typestab_tap() — track frequency distributions across
pipeline stepsaudit_diff() — before/after comparison of any two
snapshotsaudit_report() — full pipeline report in one callaudit_export() — self-contained HTML visualizationwrite_trail() / read_trail() — serialize
to RDS or JSON for CI pipelines and dashboards.numeric_summary,
.cols_include, .cols_exclude) — fine-tune what
each tap capturesStandalone functions for interactive data exploration — the questions you ask in the console before, during, and after building a pipeline.
validate_join() — analyze a join before performing it
(match rates, duplicates, unmatched keys)validate_primary_keys() /
validate_var_relationship() — key and relationship
validationcompare_tables() — column, row, numeric, and
categorical comparison between two data framesfilter_keep() / filter_drop() — filter
with diagnostic output and configurable warning thresholdsdiagnose_nas() / summarize_column() /
get_summary_table() — data quality diagnosticsdiagnose_strings() / audit_transform() —
string quality auditing and type-aware transformation diagnosticstab() — frequency tables and crosstabulations with
sorting, cutoffs, and weighting# Install from CRAN
install.packages("tidyaudit")
# Install development version
pak::pak("fpcordeiro/tidyaudit")tidyaudit is the tidyverse-native counterpart to dtaudit, a data.table-based package on CRAN. Same design vocabulary, independent implementations — choose the one that matches your stack.
LGPL (>= 3)
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.