The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
tabpfn, meaning prior fitted networks for tabular data, is a deep-learning model. See:
This R package is a wrapper of the Python library via reticulate. It has an idiomatic R syntax using standard S3 methods.
You can install the development version of tabpfn like so:
require(pak)
pak(c("tidymodels/tabpfn"), ask = FALSE)You’ll need a Python virtual environment to access the underlying library. After installing the R package, tabpfn will install the required Python bits when you first fit a model:
> library(tabpfn)
>
> predictors <- mtcars[, -1]
> outcome <- mtcars[, 1]
>
> # XY interface
> mod <- tab_pfn(predictors, outcome)
Downloading uv...Done!
Downloading cpython-3.12.12 (download) (15.9MiB)
Downloading cpython-3.12.12 (download)
Downloading setuptools (1.1MiB)
Downloading scikit-learn (8.2MiB)
Downloading numpy (4.9MiB)
<downloading and installing more packages>
Downloading llvmlite
Downloading torch
Installed 58 packages in 350ms
> mod
tabpfn Regression Model
Training set
i 32 data points
i 10 predictorslibrary(tabpfn)To fit a model:
set.seed(364)
reg_mod <- tab_pfn(mtcars[1:25, -1], mtcars$mpg[1:25])
reg_mod
#> TabPFN Regression Model
#> Training set
#> ℹ 25 data points
#> ℹ 10 predictorsIn addition to the x/y interface shown above, there are also formula and recipes interfaces.
Prediction follows the usual S3 predict() method:
predict(reg_mod, mtcars[26:32, -1])
#> # A tibble: 7 × 1
#> .pred
#> <dbl>
#> 1 29.8
#> 2 25.6
#> 3 26.2
#> 4 16.5
#> 5 19.4
#> 6 14.7
#> 7 23.6tabpfn follows the tidymodels prediction convention: a data frame is always returned with a standard set of column names.
For a classification model, the outcome should always be a factor vector. For example, using these data from the modeldata package:
library(modeldata)
#>
#> Attaching package: 'modeldata'
#> The following object is masked from 'package:datasets':
#>
#> penguins
library(ggplot2)
two_cls_train <- parabolic[1:400, ]
two_cls_val <- parabolic[401:500,]
grid <- expand.grid(X1 = seq(-5.1, 5.0, length.out = 25),
X2 = seq(-5.5, 4.0, length.out = 25))
set.seed(3824)
cls_mod <- tab_pfn(class ~ ., data = two_cls_train)
grid_pred <- predict(cls_mod, grid)
grid_pred
#> # A tibble: 625 × 3
#> .pred_Class1 .pred_Class2 .pred_class
#> <dbl> <dbl> <fct>
#> 1 0.988 0.0122 Class1
#> 2 0.992 0.00823 Class1
#> 3 0.993 0.00721 Class1
#> 4 0.993 0.00714 Class1
#> 5 0.991 0.00944 Class1
#> 6 0.982 0.0175 Class1
#> 7 0.965 0.0347 Class1
#> 8 0.922 0.0775 Class1
#> 9 0.799 0.201 Class1
#> 10 0.554 0.446 Class1
#> # ℹ 615 more rowsThe fit looks fairly good when shown with out-of-sample data:
cbind(grid, grid_pred) |>
ggplot(aes(X1, X2)) +
geom_point(data = two_cls_val, aes(col = class, pch = class),
alpha = 3 / 4, cex = 3) +
geom_contour(aes(z = .pred_Class1), breaks = 1/ 2, col = "black", linewidth = 1) +
coord_equal(ratio = 1)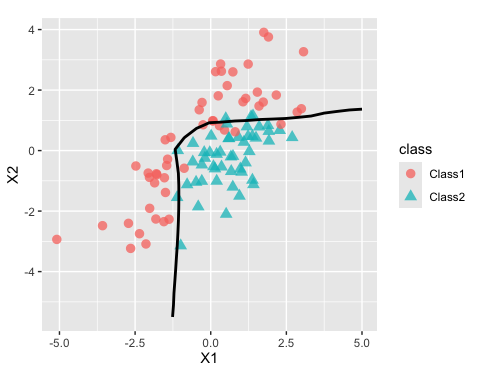
Please note that the tabpfn project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.