syntheticdata
Synthetic Clinical Data Generation with Privacy-Utility Validation
The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
Synthetic Clinical Data Generation with Privacy-Utility Validation
syntheticdata generates synthetic clinical datasets that
preserve statistical properties while reducing re-identification risk.
Useful for privacy-aware data sharing in multi-site clinical
research.
compare_methods() runs
all methods on the same data; model_fidelity() measures
train-on-synthetic, test-on-real predictive performanceUnlike synthpop (survey data) or simPop
(census microsimulation), syntheticdata integrates
generation with privacy-utility validation in a single lightweight
framework oriented toward clinical research.
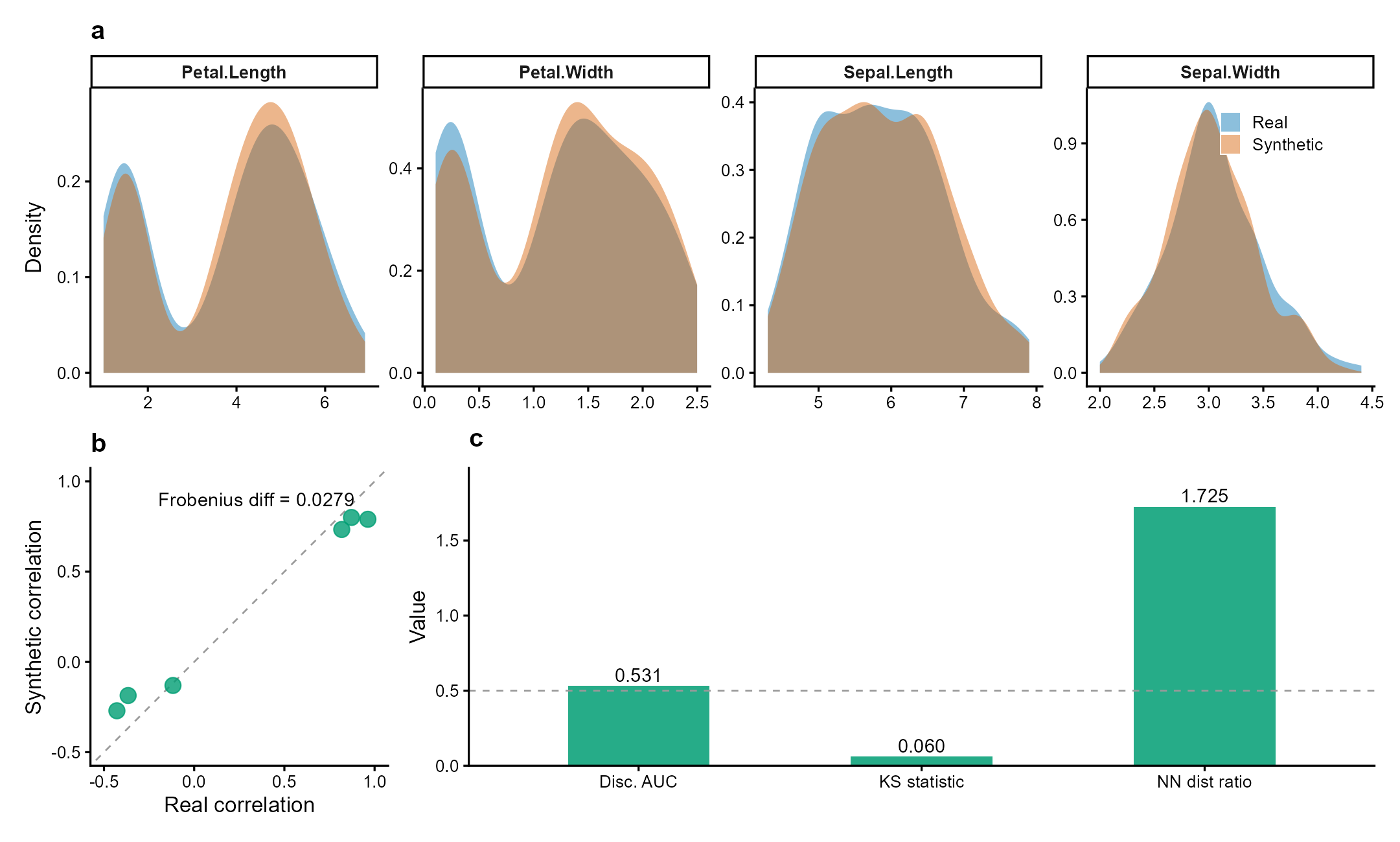
Figure 1 | Synthetic data preserves statistical properties while ensuring privacy. Fisher’s iris dataset (n = 150, 4 numeric variables) synthesized via Gaussian copula. (a) Marginal density overlays: synthetic (orange) closely matches real (blue) across all variables (mean KS = 0.06). (b) Pairwise correlation preservation (Frobenius diff = 0.028). (c) Validation metrics: discriminative AUC = 0.53 (indistinguishable from random), nearest-neighbor distance ratio = 1.73 (no privacy leakage). Data: Fisher (1936) Ann. Eugenics 7:179.
| Package | Focus | syntheticdata difference |
|---|---|---|
synthpop |
Survey/census data (CART-based) | syntheticdata targets clinical data with Gaussian copula preserving correlation structure |
simPop |
Population microsimulation | syntheticdata integrates privacy metrics (NN ratio, membership inference) |
simstudy |
Simulation for trials | syntheticdata generates from real data, not from specified distributions |
The gap: no CRAN package combines generation + privacy assessment + downstream model fidelity testing in one workflow. Existing tools either generate without validating, or validate without privacy-aware metrics.
# Complete workflow in 3 lines
syn <- synthesize(clinical_data, method = "parametric")
privacy_risk(syn, sensitive_cols = c("diagnosis", "age"))
model_fidelity(syn, outcome = "readmission")# From GitHub:
devtools::install_github("CuiweiG/syntheticdata")
# After CRAN acceptance:
install.packages("syntheticdata")library(syntheticdata)
# Synthesize from real clinical data
syn <- synthesize(iris, method = "parametric", seed = 42)
syn
# Validate utility and privacy
validate_synthetic(syn)| Function | Description |
|---|---|
synthesize() |
Generate synthetic data (parametric / bootstrap / noise) |
validate_synthetic() |
Compute utility and privacy metrics (KS, AUC, NN ratio) |
compare_methods() |
Benchmark all 3 methods on the same dataset |
privacy_risk() |
Assess re-identification risk (NN ratio, membership inference, attribute disclosure) |
model_fidelity() |
Train-on-synthetic, test-on-real predictive model comparison |
MIT
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.