The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
Website and Source code
The funStatTest package implements various statistics
for two sample comparison testing regarding functional data introduced
and used in Smida et al 2022 [1].
This package is developed by:
To install the funStatTest package, you can run:
install.packages("funStatTest")You can also install the development version of
funStatTest with the following command:
remotes::install_git("https://plmlab.math.cnrs.fr/gdurif/funStatTest")See the package vignette and function manuals for more details about the package usage.
The funStatTest was developed using the fusen
package [2]. See in the dev sub-directory in the package
sources for more information, in particular:
dev/dev_history.Rmd describing the development
processdev/flat_package.Rmd defining the major
package functions (from which the vignette is extracted)dev/flat_internal.Rmd defining package
internal functionsThe funStatTest website was generated using the pkgdown package
[3].
This is a basic example which shows you how to solve a common problem:
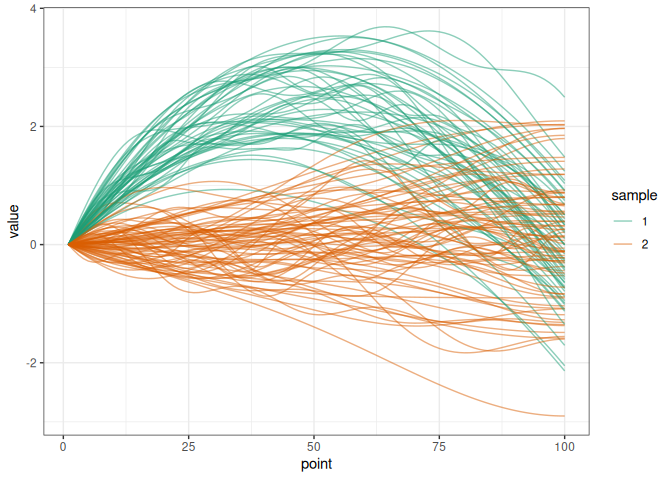
library(funStatTest)We simulate two samples of trajectories diverging by a delta function.
simu_data <- simul_data(
n_point = 100, n_obs1 = 50, n_obs2 = 75, c_val = 10,
delta_shape = "quadratic", distrib = "normal"
)
plot_simu(simu_data)
We extract the matrices of trajectories associated to each sample:
MatX <- simu_data$mat_sample1
MatY <- simu_data$mat_sample2And we compute the different statistics for two sample function data comparison presented in Smida et al 2022 [1]:
res <- comp_stat(MatX, MatY, stat = c("mo", "med", "wmw", "hkr", "cff"))
res
#> $mo
#> [1] 0.9486241
#>
#> $med
#> [1] 0.9517283
#>
#> $wmw
#> [1] 0.9074959
#>
#> $hkr
#> [,1]
#> T1 31987.663
#> T2 8489.875
#>
#> $cff
#> [1] 14150.96We can also compute p-values associated to these statistics:
# small data for the example
simu_data <- simul_data(
n_point = 20, n_obs1 = 4, n_obs2 = 5, c_val = 10,
delta_shape = "constant", distrib = "normal"
)
MatX <- simu_data$mat_sample1
MatY <- simu_data$mat_sample2
res <- permut_pval(
MatX, MatY, n_perm = 200, stat = c("mo", "med", "wmw", "hkr", "cff"),
verbose = TRUE)
res
#> $mo
#> [1] 0.01492537
#>
#> $med
#> [1] 0.0199005
#>
#> $wmw
#> [1] 0.01492537
#>
#> $hkr
#> T1 T2
#> 0.014925373 0.009950249
#>
#> $cff
#> [1] 0.009950249:warning: computing p-values based on permutations may take some time (for large data or when using a large number of simulations. :warning:
And we can also run a simulation-based power analysis:
# simulate a few small data for the example
res <- power_exp(
n_simu = 20, alpha = 0.05, n_perm = 200,
stat = c("mo", "med", "wmw", "hkr", "cff"),
n_point = 25, n_obs1 = 4, n_obs2 = 5, c_val = 10, delta_shape = "constant",
distrib = "normal", max_iter = 10000, verbose = FALSE
)
res$power_res
#> $mo
#> [1] 1
#>
#> $med
#> [1] 1
#>
#> $wmw
#> [1] 1
#>
#> $hkr
#> T1 T2
#> 1 1
#>
#> $cff
#> [1] 1These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.