The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
forrest creates publication-ready forest plots from any
data frame that contains point estimates and confidence intervals —
regression models, subgroup analyses, meta-analyses, dose-response
patterns, and more. A single dependency (tinyplot) keeps
the footprint minimal.
Key features:
section argument) — no manual NA rowssubsection for nested grouping
structuresis_summary)" (Ref.)"stripe = TRUE)section_colssave_forrest()data.frame, tibble, and
data.tableinstall.packages("forrest")
# Development version from GitHub:
# install.packages("pak")
pak::pak("lorenzoFabbri/forrest")The minimal call requires only three column names —
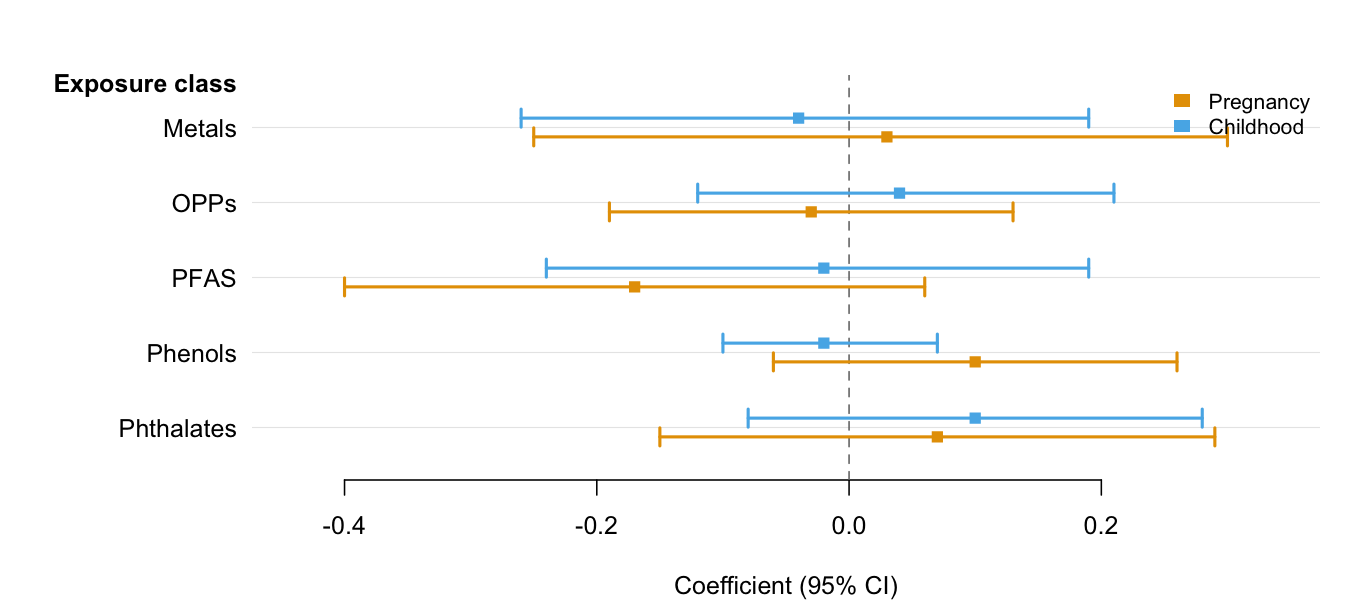
estimate, lower, and upper. Use
group and dodge = TRUE to place multiple CIs
per row when comparing results across categories (e.g., time periods,
sexes, models). See the Introduction
vignette for the full feature tour.
library(forrest)
# Two estimates per exposure (e.g., two time windows)
periods <- data.frame(
exposure = rep(c("Metals", "OPPs", "PFAS", "Phenols", "Phthalates"), each = 2),
period = rep(c("Pregnancy", "Childhood"), 5),
est = c( 0.03, -0.04, -0.03, 0.04, -0.17, -0.02, 0.10, -0.02, 0.07, 0.10),
lo = c(-0.25, -0.26, -0.19, -0.12, -0.40, -0.24, -0.06, -0.10, -0.15, -0.08),
hi = c( 0.30, 0.19, 0.13, 0.21, 0.06, 0.19, 0.26, 0.07, 0.29, 0.28)
)
forrest(
periods,
estimate = "est",
lower = "lo",
upper = "hi",
label = "exposure",
group = "period",
dodge = TRUE,
header = "Exposure class",
ref_line = 0,
xlab = "Coefficient (95% CI)"
)
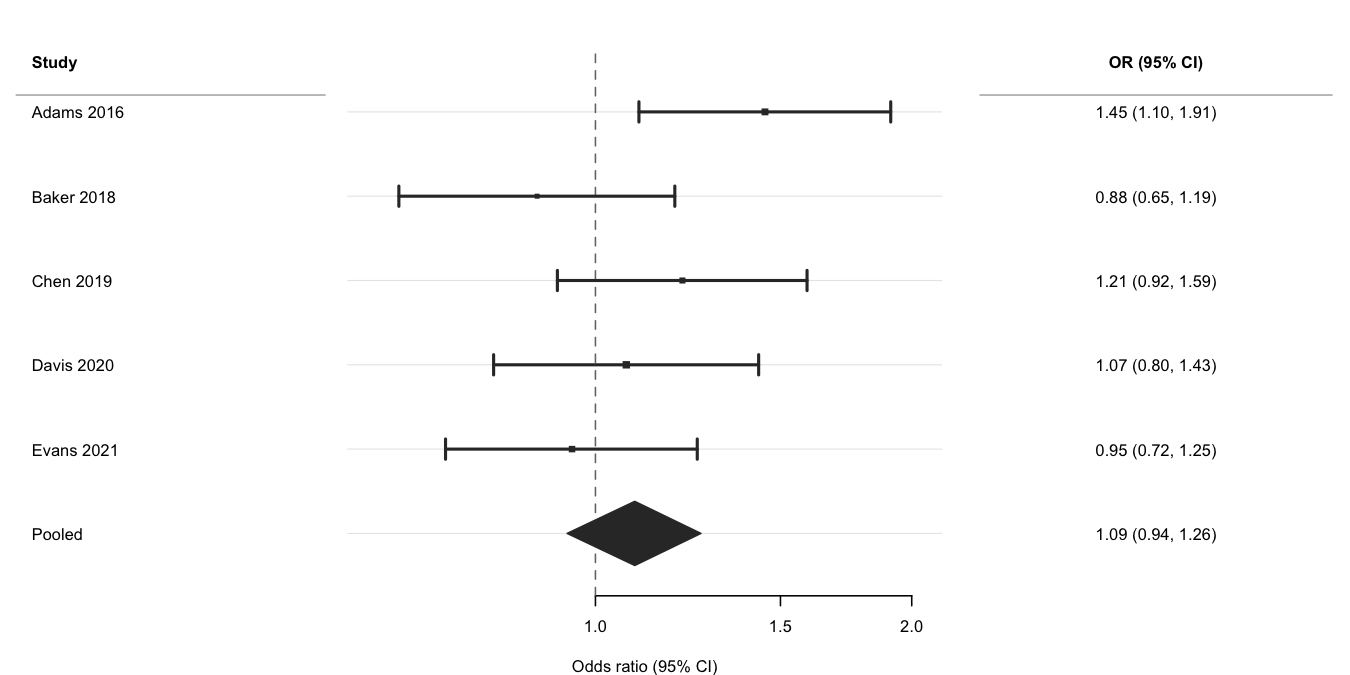
Use log_scale = TRUE and ref_line = 1 for
ratio measures, is_summary = TRUE to draw the pooled row as
a filled diamond, and weight to scale point sizes by
inverse-variance weights. See the Meta-analysis
vignette for subgroup analyses and multi-exposure layouts.
ma <- data.frame(
study = c("Adams 2016", "Baker 2018", "Chen 2019",
"Davis 2020", "Evans 2021", "Pooled"),
est = c(1.45, 0.88, 1.21, 1.07, 0.95, 1.09),
lo = c(1.10, 0.65, 0.92, 0.80, 0.72, 0.94),
hi = c(1.91, 1.19, 1.59, 1.43, 1.25, 1.26),
weight = c(180, 95, 140, 210, 160, NA),
is_sum = c(FALSE, FALSE, FALSE, FALSE, FALSE, TRUE)
)
ma$or_ci <- sprintf("%.2f (%.2f, %.2f)", ma$est, ma$lo, ma$hi)
forrest(
ma,
estimate = "est",
lower = "lo",
upper = "hi",
label = "study",
is_summary = "is_sum",
weight = "weight",
log_scale = TRUE,
ref_line = 1,
header = "Study",
cols = c("OR (95% CI)" = "or_ci"),
widths = c(2.2, 4, 2.5),
xlab = "Odds ratio (95% CI)"
)
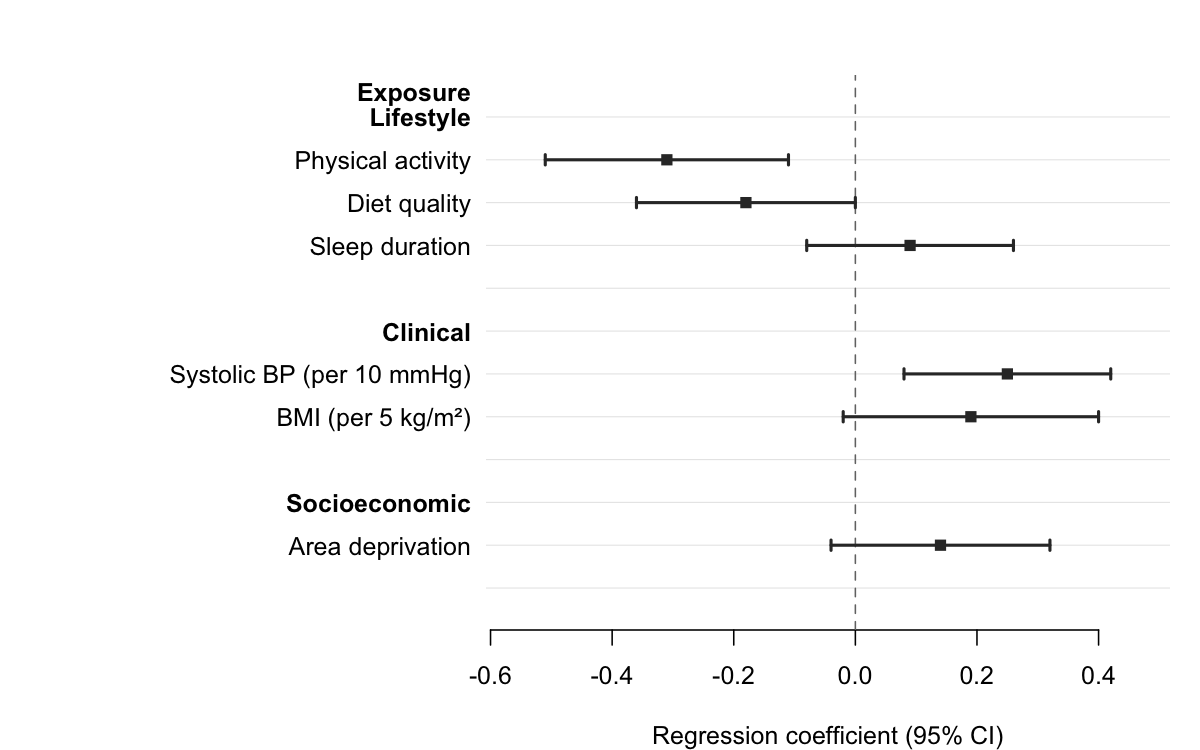
Pass a column name to section to group rows under
automatic bold headers. forrest() inserts a header wherever
the section value changes, indents the row labels, and adds a blank
spacer after each group — no manual data manipulation required.
subgroups <- data.frame(
domain = c("Lifestyle", "Lifestyle", "Lifestyle",
"Clinical", "Clinical",
"Socioeconomic"),
exposure = c("Physical activity", "Diet quality", "Sleep duration",
"Systolic BP (per 10 mmHg)", "BMI (per 5 kg/m\u00b2)",
"Area deprivation"),
est = c(-0.31, -0.18, 0.09, 0.25, 0.19, 0.14),
lo = c(-0.51, -0.36, -0.08, 0.08, -0.02, -0.04),
hi = c(-0.11, -0.00, 0.26, 0.42, 0.40, 0.32)
)
forrest(
subgroups,
estimate = "est",
lower = "lo",
upper = "hi",
label = "exposure",
section = "domain",
header = "Exposure",
ref_line = 0,
xlab = "Regression coefficient (95% CI)"
)
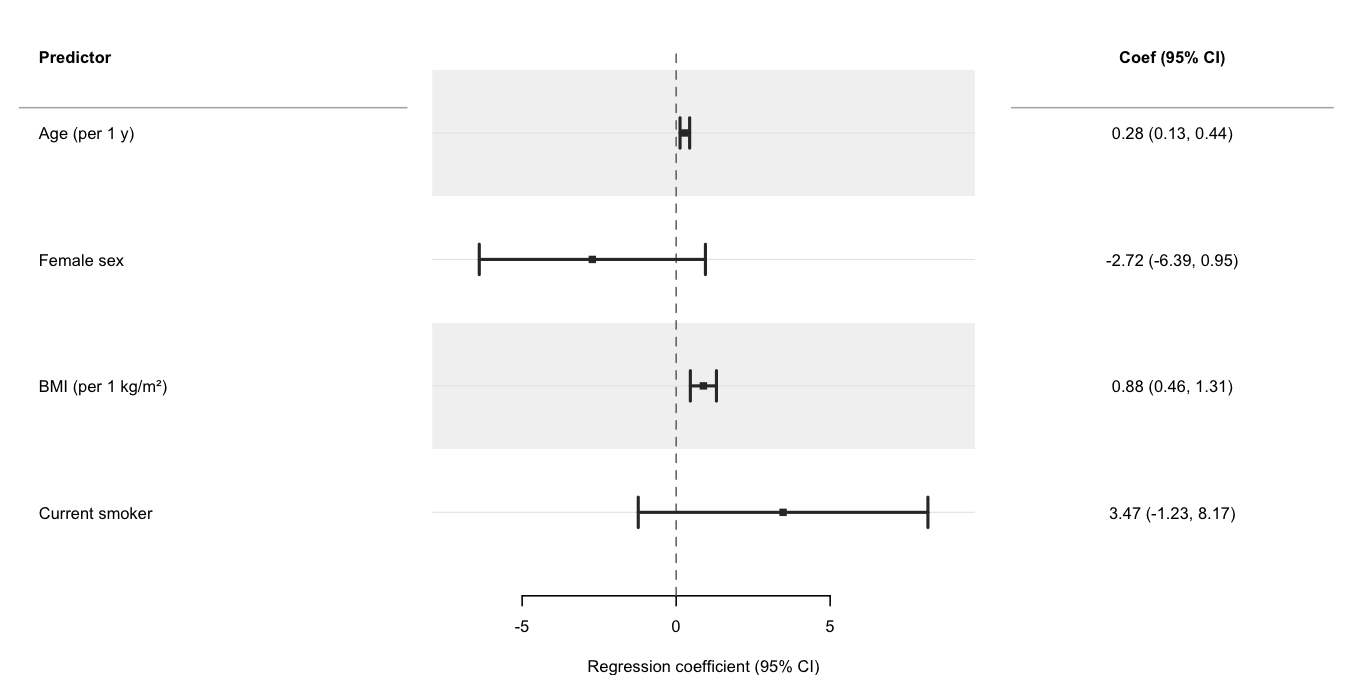
Use broom::tidy() to
extract parameters from a fitted model and pass the result directly to
forrest(). The cols argument adds a formatted
text column alongside the plot; header labels the row-label
panel. See the Regression
models vignette for progressive adjustment, multiple outcomes,
dose-response, and causal inference (g-computation, IPW) examples.
set.seed(1)
n <- 300
dat <- data.frame(
sbp = 120 + rnorm(n, sd = 15),
age = runif(n, 30, 70),
female = rbinom(n, 1, 0.5),
bmi = rnorm(n, 26, 4),
smoker = rbinom(n, 1, 0.2)
)
dat$sbp <- dat$sbp + 0.3 * dat$age - 4 * dat$female +
0.5 * dat$bmi - 2 * dat$smoker + rnorm(n, sd = 6)
fit <- lm(sbp ~ age + female + bmi + smoker, data = dat)
coefs <- broom::tidy(fit, conf.int = TRUE)
coefs <- coefs[coefs$term != "(Intercept)", ]
coefs$term <- c("Age (per 1 y)", "Female sex",
"BMI (per 1 kg/m²)", "Current smoker")
coefs$coef_ci <- sprintf(
"%.2f (%.2f, %.2f)",
coefs$estimate, coefs$conf.low, coefs$conf.high
)
forrest(
coefs,
estimate = "estimate",
lower = "conf.low",
upper = "conf.high",
label = "term",
header = "Predictor",
cols = c("Coef (95% CI)" = "coef_ci"),
widths = c(3, 4, 2.5),
xlab = "Regression coefficient (95% CI)",
stripe = TRUE
)
| Feature | forrest | forestplot | forestploter | ggforestplot |
|---|---|---|---|---|
| Minimal dependencies | ✅ | ❌ | ❌ | ❌ |
| General (not meta-analysis only) | ✅ | ✅ | ✅ | ⚠️ |
| Study weight sizing | ✅ | ✅ | ✅ | ❌ |
| Auto section headers from grouping column | ✅ | ⚠️ | ❌ | ❌ |
| Two-level nested section hierarchy | ✅ | ❌ | ❌ | ❌ |
| Summary diamonds | ✅ | ✅ | ✅ | ❌ |
| Text columns | ✅ | ✅ | ✅ | ❌ |
| Alternating row stripes | ✅ | ✅ | ✅ | ✅ |
| CI clipping with arrows | ✅ | ✅ | ⚠️ | ❌ |
| Group colouring + legend | ✅ | ✅ | ✅ | ✅ |
| Log-scale axis | ✅ | ✅ | ✅ | ✅ |
| Export helper | ✅ | ❌ | ⚠️ | ❌ |
| data.table support | ✅ | ❌ | ❌ | ❌ |
| Actively maintained | ✅ | ✅ | ✅ | ❌ |
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.