The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
Dynamic Failure Rate Distributions for Survival Analysis
Capacitors that wear out faster than any Weibull can describe.
Software systems with bathtub-shaped crash rates. Post-surgical patients
whose risk drops sharply, then slowly climbs again. Standard parametric
survival families cannot express these hazard patterns — but
flexhaz can.
Write the hazard function you need — any R function of time and parameters — and the package derives everything else: survival curves, CDFs, densities, quantiles, sampling, log-likelihoods, MLE fitting, and residual diagnostics.
| Feature | flexhaz | survival | flexsurv |
|---|---|---|---|
| Custom hazard functions | Yes | No | Limited |
| Built-in distributions | Exp, Weibull, Gompertz, Log-logistic | Weibull, Exp | Many |
| User-supplied derivatives | score + Hessian | No | No |
| Censoring support | Right + Left | Right | Right |
| Model diagnostics | Cox-Snell, Martingale, Q-Q | Limited | Limited |
| Likelihood model interface | Full | Basic | Partial |
algebraic.dist, likelihood.model,
algebraic.mleInstall from CRAN:
install.packages("flexhaz")Or the development version from r-universe:
install.packages("flexhaz", repos = "https://queelius.r-universe.dev")library(flexhaz)Use the convenient constructors for classic survival distributions:
# Exponential: constant hazard (memoryless)
exp_dist <- dfr_exponential(lambda = 0.5)
# Weibull: power-law hazard (wear-out or infant mortality)
weib_dist <- dfr_weibull(shape = 2, scale = 3)
# Gompertz: exponentially increasing hazard (aging)
gomp_dist <- dfr_gompertz(a = 0.01, b = 0.1)
# Log-logistic: non-monotonic hazard (increases then decreases)
ll_dist <- dfr_loglogistic(alpha = 10, beta = 2)All distribution functions are automatically available:
S <- surv(exp_dist)
S(2) # Survival probability at t=2
#> [1] 0.3678794
h <- hazard(weib_dist)
h(1) # Hazard at t=1
#> [1] 0.2222222# Simulate failure times
set.seed(42)
times <- rexp(50, rate = 1)
df <- data.frame(t = times, delta = 1)
# Fit via MLE
solver <- fit(dfr_exponential())
result <- solver(df, par = c(0.5), method = "BFGS")
coef(result) # Estimated rate
#> [1] 0.8808457Model complex failure patterns like bathtub curves:
# h(t) = a*exp(-b*t) + c + d*t^k
# Infant mortality + useful life + wear-out
bathtub <- dfr_dist(
rate = function(t, par, ...) {
par[1] * exp(-par[2] * t) + par[3] + par[4] * t^par[5]
},
par = c(a = 1, b = 2, c = 0.02, d = 0.001, k = 2)
)
h <- hazard(bathtub)
curve(sapply(x, h), 0, 15, xlab = "Time", ylab = "Hazard rate",
main = "Bathtub hazard curve")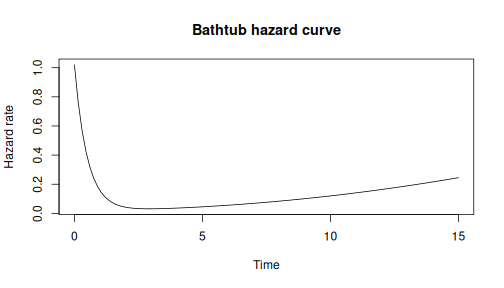
Check model fit with residual analysis:
# Fit exponential to data
fitted_exp <- dfr_exponential(lambda = coef(result))
# Cox-Snell residuals Q-Q plot
qqplot_residuals(fitted_exp, df)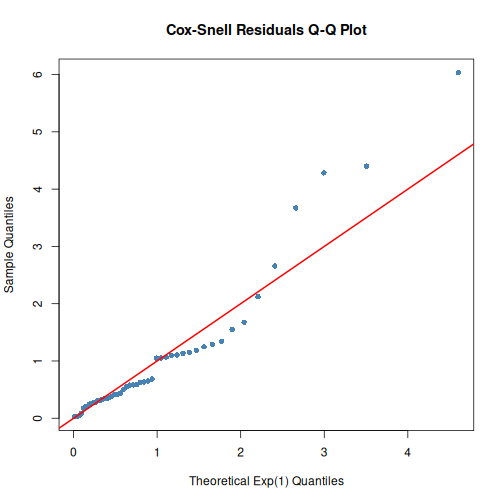
For a lifetime , the hazard function is:
From the hazard, all other quantities follow:
| Function | Formula | Method |
|---|---|---|
| Cumulative hazard | cum_haz() |
|
| Survival | surv() |
|
| CDF | cdf() |
|
density() |
For exact observations:
For right-censored:
# Mixed data with censoring
df <- data.frame(
t = c(1, 2, 3, 4, 5),
delta = c(1, 1, 0, 1, 0) # 1 = exact, 0 = censored
)
ll <- loglik(dfr_exponential())
ll(df, par = c(0.5))
#> [1] -9.579442Start Here:
Real-World Applications:
Going Deeper:
Reference:
algebraic.dist:
Generic distribution interfacelikelihood.model:
Likelihood model frameworkalgebraic.mle:
MLE utilitiesThese binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.