clinicalfair
Algorithmic Fairness Assessment for Clinical Prediction Models
The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.
Algorithmic Fairness Assessment for Clinical Prediction Models
clinicalfair is a post-hoc fairness auditing toolkit for
clinical prediction models. It evaluates existing models by computing
group-wise fairness metrics, visualizing disparities, and performing
threshold-based mitigation – motivated by regulatory expectations for
transparency in clinical AI.
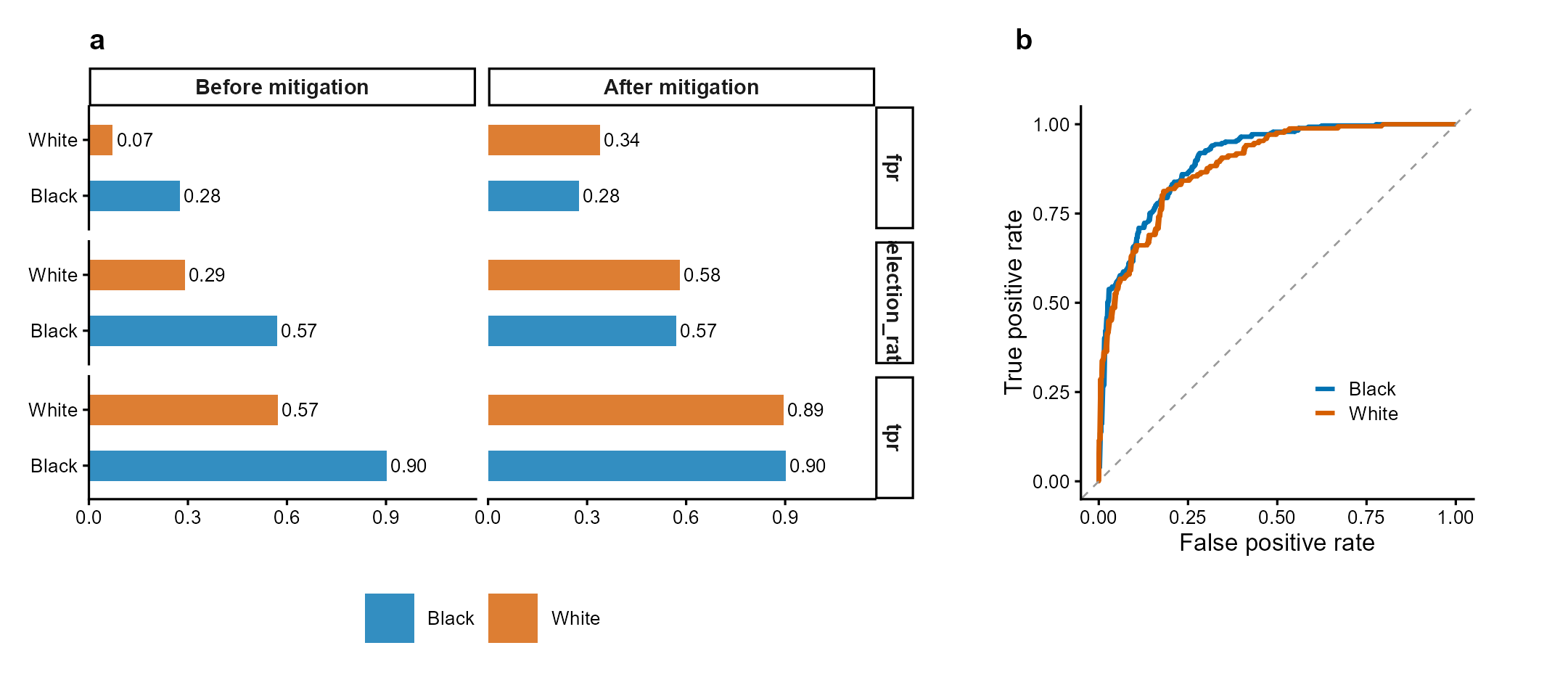
Figure 1 | Fairness audit with threshold mitigation. (a) Group-wise selection rate, TPR, and FPR before and after equalized-odds threshold optimization. Mitigation reduces cross-group TPR disparity while maintaining acceptable accuracy. (b) ROC curves by racial group showing differential model performance. Data: COMPAS-style simulated recidivism predictions. Methods: Hardt et al. (2016); Obermeyer et al. (2019).
Existing R packages approach fairness from different angles:
| Package | Focus | clinicalfair difference |
|---|---|---|
fairmodels |
Model-level fairness (wraps mlr3) | clinicalfair is model-agnostic: works with any predicted probabilities |
fairness |
Metric computation | clinicalfair adds threshold mitigation + intersectional analysis |
fairmetrics |
CIs for fairness metrics | clinicalfair also provides bootstrap CIs, plus audit reports with four-fifths rule screening |
clinicalfair is designed for the clinical audit
use case: a clinician or regulator receives a trained model and
needs to answer “is this model fair across patient subgroups?” in one
function call, with actionable output.
# One-line audit
fairness_report(fairness_data(predictions, labels, race))# From GitHub:
devtools::install_github("CuiweiG/clinicalfair")
# After CRAN acceptance:
install.packages("clinicalfair")library(clinicalfair)
data(compas_sim)
# Create fairness evaluation object
fd <- fairness_data(
predictions = compas_sim$risk_score,
labels = compas_sim$recidivism,
protected_attr = compas_sim$race
)
# Compute metrics
fairness_metrics(fd)
# With bootstrap confidence intervals
fairness_metrics(fd, ci = TRUE, n_boot = 2000)
# Generate audit report
fairness_report(fd)
# Mitigate via threshold optimization
threshold_optimize(fd, objective = "equalized_odds")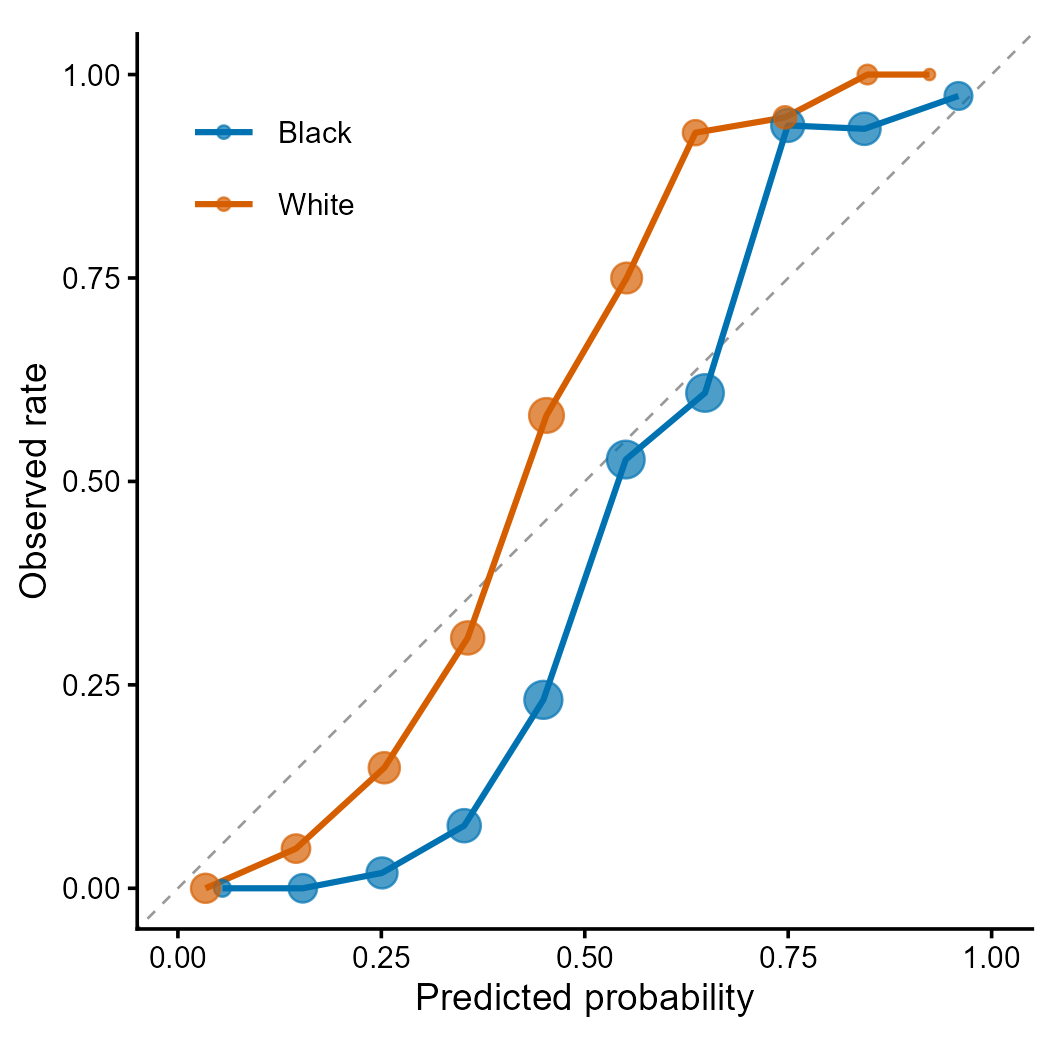
Figure 2 | Calibration curves by racial group. Each point represents a decile bin; point size proportional to sample count. Deviation from the diagonal indicates miscalibration. Differential calibration across groups is a key fairness concern identified by Obermeyer et al. (2019) Science 366:447.
| Function | Description |
|---|---|
fairness_data() |
Bundle predictions, labels, protected attributes |
fairness_metrics() |
Compute group-wise fairness metrics (optional bootstrap CIs) |
fairness_report() |
Generate audit report with four-fifths rule screening |
threshold_optimize() |
Group-specific threshold mitigation (configurable grid resolution) |
intersectional_fairness() |
Cross-tabulated multi-attribute analysis (small-group filtering) |
autoplot() |
Disparity bar plots (S3 method) |
plot_roc() |
ROC curves by protected group |
plot_calibration() |
Calibration curves by group |
MIT
These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.