
The hardware and bandwidth for this mirror is donated by dogado GmbH, the Webhosting and Full Service-Cloud Provider. Check out our Wordpress Tutorial.
If you wish to report a bug, or if you are interested in having us mirror your free-software or open-source project, please feel free to contact us at mirror[@]dogado.de.

RSQLite.toolkit is a lightweight wrapper around the RSQLite package for streamlined loading of data from tabular files (i.e. text delimited files like CSV and TSV, Microsoft Excel, and Arrow IPC files) in SQLite databases.
It also includes helper functions for inspecting the structure of the input files, and some functions to simplify activities on the SQLite tables.
Install the released version of RSQLite.toolkit from CRAN:
install.packages("RSQLite.toolkit")You can install the development version from GitHub with:
# Install development version from GitHub
# install.packages("pak")
pak::pak("fab-algo/RSQLite.toolkit")These basic examples show how to use the core functions of the package to load example data in different tables in a test database:
library(RSQLite.toolkit)
dbcon <- dbConnect(RSQLite::SQLite(), file.path(tempdir(), "tests.sqlite"))
data_path <- system.file("extdata", package = "RSQLite.toolkit")
## creates the table ABALONE from a CSV file, adding the field 'SEQ' with the
## ROWID of each record through the use of the parameter 'auto_pk=TRUE'
dbTableFromDSV(input_file = file.path(data_path, "abalone.csv"),
dbcon = dbcon, table_name = "ABALONE",
drop_table = TRUE, auto_pk = TRUE,
header = TRUE, sep = ",", dec = ".")
#> [1] 4177
## creates the table PORTFOLIO_PERF from Excel file, using the "all period" sheet
dbTableFromXlsx(input_file = file.path(data_path,
"stock_portfolio.xlsx"),
dbcon = dbcon, table_name = "PORTFOLIO_PERF",
drop_table = TRUE,
sheet_name = "all period", first_row = 2, cols_range = "A:S")
#> [1] 63
## creates the table PENGUINS from Feather file
dbTableFromFeather(input_file = file.path(data_path, "penguins.feather"),
dbcon = dbcon, table_name = "PENGUINS",
drop_table = TRUE)
#> [1] 333
dbListTables(dbcon)
#> [1] "ABALONE" "PENGUINS" "PORTFOLIO_PERF"
dbListFields(dbcon, "ABALONE")
#> [1] "Sex" "Length" "Diam" "Height" "Whole" "Shucked" "Viscera"
#> [8] "Shell" "Rings" "SEQ"
dbListFields(dbcon, "PENGUINS")
#> [1] "species" "culmen_length_mm" "culmen_depth_mm"
#> [4] "flipper_length_mm" "body_mass_g" "sex"
dbListFields(dbcon, "PORTFOLIO_PERF")[1:5]
#> [1] "ID"
#> [2] "Large_B_P"
#> [3] "Large_ROE"
#> [4] "Large_S_P"
#> [5] "Large_Return_Rate_in_the_last_quarter"
dbDisconnect(dbcon)The basic idea behind this package is that storing all the data used throughout a data-analysis workflow in a local database is a highly efficient and effective way to manage information. The advantages of keeping raw data and their metadata together in the same database—ideally under a shared data model—far outweigh not only the additional overhead of maintaining a structured system, but also the costs and complexity involved in importing information from external files with disparate formats and conventions.
The purpose of this package is to reduce the burden of the last point (i.e., importing the data) as much as possible through a set of functions that read the data from the source files, create the destination table (if it does not exist), store the data there, and index it, all in one step.
The core functions of the package are therefore those that can be used to move data from a file to a database table:
dbTableFromDSV()dbTableFromXlsx()dbTableFromFeather()Together with these, there are a couple of additional functions:
dbTableFromDataFrame()dbTableFromView()that could be handy to move data from intermediate data structures to database tables, using the same logic of the core ones.
All these functions share a common calling template that is outlined in the following picture:
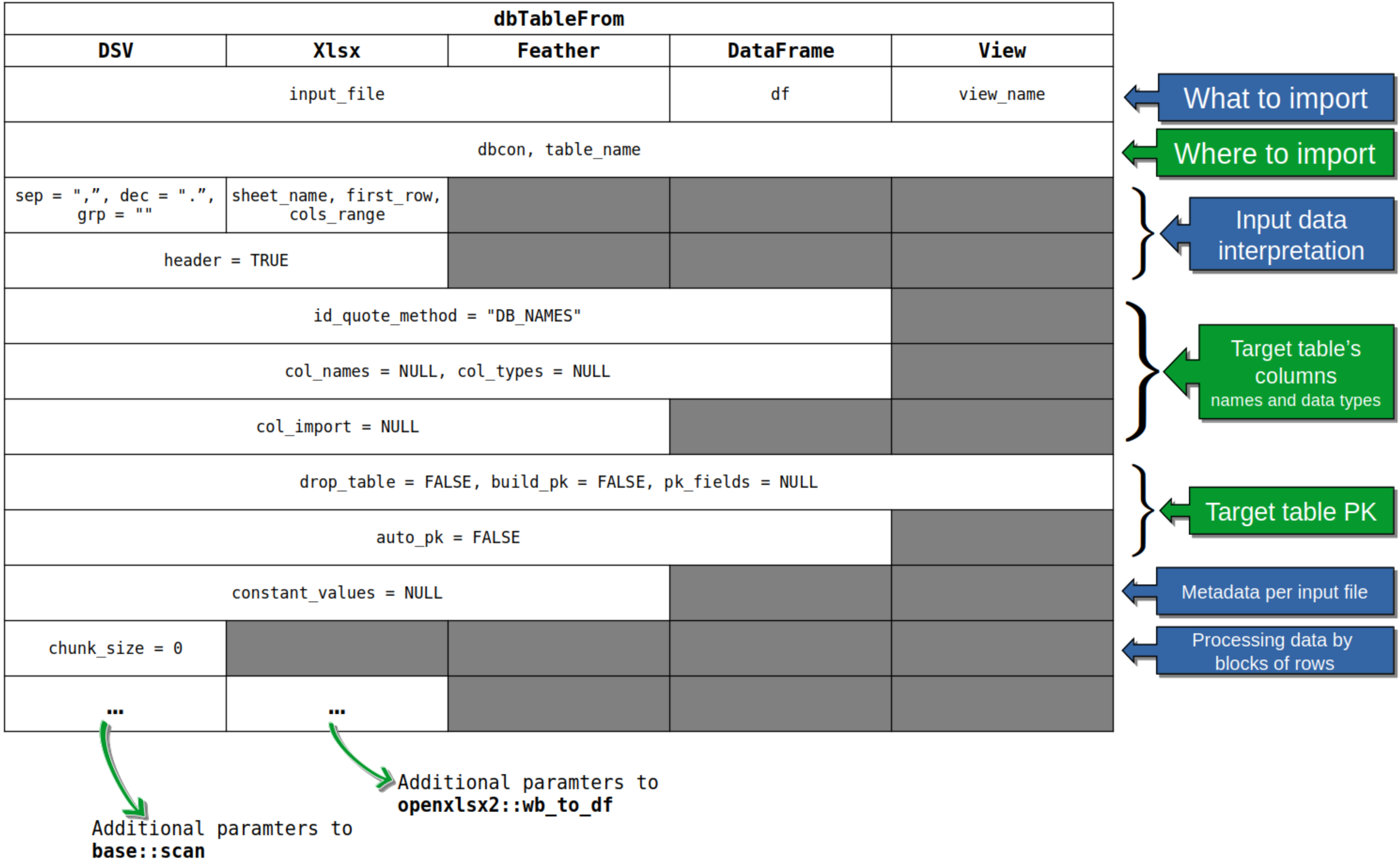
All core functions require three mandatory arguments:
input_filedbcontable_namethat have no default values. The dbTableFromXlsx()
requires three additional arguments (again with no defaults):
sheet_name, first_row, cols_range
that are needed to correctly locate the dataset to be imported inside
the Excel file.
More details on the parameters and functions’ behaviour are available in the standard function documentation accessible through R’s help system.
Anyway, be careful with default values of all other arguments, especially when importing from delimiter separated values text files: the default values for input data interpretation arguments are seldom the right ones. This is also true for the default values of the functions actually used by this package to read data from files, that is:
base::scan() for reading from DSV files;openxlsx2::wb_to_df() for reading from Xlsx files;arrow::FeatherReader() for reading from Feather
files.Given the high variability in how data can be stored in DSV and Excel
files, there is always the possibility of passing specific parameters of
the above-listed functions through the special argument
“...”. To help with using the additional arguments of
dbTableFromDSV(), along with those of
base::scan(), you can read the vignette “Dealing
with DSV files” that is part of this package. The first section
of the document describes how to use base::scan() arguments
to read complex DSV files. In the second section there is a detailed
description of the issues related to the column names used in the data
file and their corresponding identifiers in the SQLite table’s fields.
The second section of this vignette can be helpful also when importing
data from Xlsx and Feather files.
Before loading data into your SQLite database, RSQLite.toolkit provides specialized functions to inspect the structure and schema of your input files without loading the entire dataset into memory. This is particularly useful for large files or when you need to understand the data types and column structure before importing.
The package includes three schema inspection functions:
file_schema_dsv() - for
delimiter-separated values files (CSV, TSV, etc.)file_schema_feather() - for Arrow
Feather filesfile_schema_xlsx() - for Microsoft
Excel filesThese functions serve several important purposes:
dbTableFrom*() functionsAll three functions return a data frame with the following key columns:
col_names: Column names after applying SQLite
formatting rulescol_names_unquoted: Unquoted column names (if different
from col_names)col_types: Inferred R data types for each columnsql_types: Corresponding SQLite data typessrc_names: Original column names as they appear in the
source filesrc_types: Source data types (varies by file
format)library(RSQLite.toolkit)
data_path <- system.file("extdata", package = "RSQLite.toolkit")
file_schema_dsv(input_file = file.path(data_path, "abalone.csv"),
header = TRUE, sep = ",", dec = ".")$schema[, c(1, 3:7)]
#> col_names col_types sql_types src_names src_types src_is_quoted
#> 1 Sex character TEXT Sex text FALSE
#> 2 Length numeric REAL Length text FALSE
#> 3 Diam numeric REAL Diam text FALSE
#> 4 Height numeric REAL Height text FALSE
#> 5 Whole numeric REAL Whole text FALSE
#> 6 Shucked numeric REAL Shucked text FALSE
#> 7 Viscera numeric REAL Viscera text FALSE
#> 8 Shell numeric REAL Shell text FALSE
#> 9 Rings integer INTEGER Rings text FALSE
file_schema_feather(input_file = file.path(data_path,
"penguins.feather"))[, c(1, 3:6)]
#> col_names col_types sql_types src_names src_types
#> 1 species character TEXT species utf8
#> 2 culmen_length_mm double REAL culmen_length_mm double
#> 3 culmen_depth_mm double REAL culmen_depth_mm double
#> 4 flipper_length_mm double REAL flipper_length_mm double
#> 5 body_mass_g double REAL body_mass_g double
#> 6 sex character TEXT sex utf8These schema functions use the same parameters as their
corresponding dbTableFrom*() functions, making it easy to
test and refine your import settings before committing to the full data
load.
Beyond the core data import capabilities, RSQLite.toolkit provides two utility functions for simplifying database operations:
dbExecFile() - Execute SQL statements
from text filesdbCopyTable() - Copy tables between
SQLite databasesThe dbExecFile() function allows you to execute multiple
SQL statements stored in a text file, making it ideal for:
Moreover, the possibility to read SQL statements directly from a text file avoids the need to embed long SQL instructions in strings inside the main R code, making the source more readable.
--);)library(RSQLite.toolkit)
dbcon <- dbConnect(RSQLite::SQLite(), file.path(tempdir(), "tests.sqlite"))
## Load data from a DSV file into a database table ------------
data_path <- system.file("extdata", package = "RSQLite.toolkit")
dbTableFromDSV(input_file = file.path(data_path, "abalone.csv"),
dbcon = dbcon, table_name = "ABALONE",
drop_table = TRUE, auto_pk = TRUE,
header = TRUE, sep = ",", dec = ".")
#> [1] 4177
## ------------------------------------------------------------
## SQL file: each statement is separated by a semicolon -------
sql_text <- paste("DROP TABLE IF EXISTS AVG_LEN_BY_SEX_RINGS;",
" ",
"CREATE TABLE AVG_LEN_BY_SEX_RINGS ",
"(SEX TEXT, RINGS INTEGER, NUM INTEGER, AVG_LENGTH REAL);",
" ",
"INSERT INTO AVG_LEN_BY_SEX_RINGS ",
"SELECT SEX, RINGS, COUNT(*) AS NUM, AVG(LENGTH) ",
"AS AVG_LENGTH FROM ABALONE GROUP BY SEX, RINGS;",
" ",
"SELECT * FROM AVG_LEN_BY_SEX_RINGS;",
" ",
"SELECT SEX, round(WHOLE/:bin_size,0)*:bin_size as WHOLE_GRP, ",
"COUNT(*) AS NUM, AVG(LENGTH) AS AVG_LENGTH ",
"FROM ABALONE GROUP BY SEX, round(WHOLE/:bin_size,0)*:bin_size;",
sep = "\n")
sql_file <- tempfile(fileext = ".sql")
writeLines(sql_text, con = sql_file)
## ------------------------------------------------------------
## Execute SQL statements from the file -----------------------
plist=list(q1=NULL,
q2=NULL,
q3=NULL,
q4=NULL,
q5=list(bin_size = 0.025))
res <- dbExecFile(input_file = sql_file, dbcon = dbcon,
plist = plist)
## ------------------------------------------------------------
## Check results ----------------------------------------------
dbListTables(dbcon)
#> [1] "ABALONE" "AVG_LEN_BY_SEX_RINGS" "PENGUINS"
#> [4] "PORTFOLIO_PERF"
dbListFields(dbcon, "AVG_LEN_BY_SEX_RINGS")
#> [1] "SEX" "RINGS" "NUM" "AVG_LENGTH"
res[[1]]
#> NULL
res[[2]]
#> NULL
res[[3]]
#> NULL
res[[4]][1:10, ]
#> SEX RINGS NUM AVG_LENGTH
#> 1 F 5 4 0.3237500
#> 2 F 6 16 0.4628125
#> 3 F 7 44 0.4678409
#> 4 F 8 122 0.5380328
#> 5 F 9 238 0.5746008
#> 6 F 10 248 0.5822782
#> 7 F 11 200 0.6137000
#> 8 F 12 128 0.5949219
#> 9 F 13 88 0.5814773
#> 10 F 14 56 0.5960714
res[[5]][1:10, ]
#> Sex WHOLE_GRP NUM AVG_LENGTH
#> 1 F 0.075 1 0.2750000
#> 2 F 0.150 3 0.3000000
#> 3 F 0.175 2 0.3425000
#> 4 F 0.200 9 0.3444444
#> 5 F 0.225 5 0.3730000
#> 6 F 0.250 3 0.3733333
#> 7 F 0.275 10 0.3700000
#> 8 F 0.300 5 0.3900000
#> 9 F 0.325 7 0.4000000
#> 10 F 0.350 9 0.4038889
## ------------------------------------------------------------
## Cleanup ----------------------------------------------------
unlink(sql_file)
dbDisconnect(dbcon)The dbCopyTable() function enables you to copy entire
tables between different SQLite database files, useful for:
The typical scenario where this function becomes really useful is when you want to start a new analysis workflow in a new environment and you want to carry over part of the data you used in a previous one. This function allows you to move whole tables from the SQLite database you used in the old workflow to the new database, so that you can add more recent data or you can reuse previous results.
# Copy a table from one database to another
dbCopyTable(
db_file_src = "source.sqlite",
db_file_tgt = "backup.sqlite",
table_name = "sales_data",
drop_table = TRUE, # Recreate table if it exists
copy_indexes = TRUE # Copy indexes too
)
# Append data to existing table (no index copying)
dbCopyTable(
db_file_src = "daily_data.sqlite",
db_file_tgt = "master.sqlite",
table_name = "transactions",
drop_table = FALSE # Append to existing table
)
# Archive old data
dbCopyTable(
db_file_src = "production.sqlite",
db_file_tgt = "archive_2024.sqlite",
table_name = "historical_sales",
drop_table = TRUE,
copy_indexes = TRUE
)Use dbExecFile() when:
Use dbCopyTable() when:
These utility functions complement the core data import functions by providing database management capabilities that are often needed in real-world data analysis workflows.
The example datasets, included in the extdata package
directory, have been retrieved from the following sources:
"abalone.csv": Predicting the age of abalone from
physical measurements. Source link: https://archive.ics.uci.edu/dataset/1/abalone"stock_portfolio.xlsx": Stock portfolio performance
under a new weighted scoring stock selection model. Source link: https://archive.ics.uci.edu/dataset/390/stock+portfolio+performance"penguins.feather": The Palmer Archipelago’s penguins
data set. Source link: https://allisonhorst.github.io/palmerpenguins/. The
“feather” file format of the dataset was downloaded from https://github.com/lmassaron/datasets.These binaries (installable software) and packages are in development.
They may not be fully stable and should be used with caution. We make no claims about them.
Health stats visible at Monitor.